Metformin exerts multitarget antileukemia activity in JAK2V617F-positive myeloproliferative neoplasms
- PMID: 29472557
- PMCID: PMC5833553
- DOI: 10.1038/s41419-017-0256-4
Metformin exerts multitarget antileukemia activity in JAK2V617F-positive myeloproliferative neoplasms
Abstract
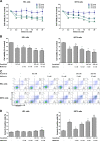
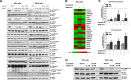
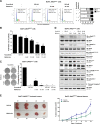
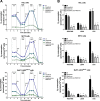
The recurrent gain-of-function JAK2V617F mutation confers growth factor-independent proliferation for hematopoietic cells and is a major contributor to the pathogenesis of myeloproliferative neoplasms (MPN). The lack of complete response in most patients treated with the JAK1/2 inhibitor ruxolitinib indicates the need for identifying novel therapeutic strategies. Metformin is a biguanide that exerts selective antineoplastic activity in hematological malignancies. In the present study, we investigate and compare effects of metformin and ruxolitinib alone and in combination on cell signaling and cellular functions in JAK2V617F-positive cells. In JAK2V617F-expressing cell lines, metformin treatment significantly reduced cell viability, cell proliferation, clonogenicity, and cellular oxygen consumption and delayed cell cycle progression. Metformin reduced cyclin D1 expression and RB, STAT3, STAT5, ERK1/2 and p70S6K phosphorylation. Metformin plus ruxolitinib demonstrated more intense reduction of cell viability and induction of apoptosis compared to monotherapy. Notably, metformin reduced Ba/F3 JAK2V617F tumor burden and splenomegaly in Jak2V617F knock-in-induced MPN mice and spontaneous erythroid colony formation in primary cells from polycythemia vera patients. In conclusion, metformin exerts multitarget antileukemia activity in MPN: downregulation of JAK2/STAT signaling and mitochondrial activity. Our exploratory study establishes novel molecular mechanisms of metformin and ruxolitinib action and provides insights for development of alternative/complementary therapeutic strategies for MPN.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


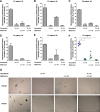
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

