Anti-HER2 scFv-Directed Extracellular Vesicle-Mediated mRNA-Based Gene Delivery Inhibits Growth of HER2-Positive Human Breast Tumor Xenografts by Prodrug Activation
- PMID: 29483213
- PMCID: PMC5932266
- DOI: 10.1158/1535-7163.MCT-17-0827
Anti-HER2 scFv-Directed Extracellular Vesicle-Mediated mRNA-Based Gene Delivery Inhibits Growth of HER2-Positive Human Breast Tumor Xenografts by Prodrug Activation
Abstract
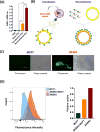
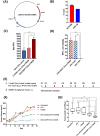
This paper deals with specific targeting of the prodrug/enzyme regimen, CNOB/HChrR6, to treat a serious disease, namely HER2+ human breast cancer with minimal off-target toxicity. HChrR6 is an improved bacterial enzyme that converts CNOB into the cytotoxic drug MCHB. Extracellular vesicles (EV) were used for mRNA-based HchrR6 gene delivery: EVs may cause minimal immune rejection, and mRNA may be superior to DNA for gene delivery. To confine HChrR6 generation and CNOB activation to the cancer, the EVHB chimeric protein was constructed. It contains high-affinity anti-HER2 scFv antibody (ML39) and is capable of latching on to EV surface. Cells transfected with EVHB-encoding plasmid generated EVs displaying this protein ("directed EVs"). Transfection of a separate batch of cells with the new plasmid, XPort/HChrR6, generated EVs containing HChrR6 mRNA; incubation with pure EVHB enabled these to target the HER2 receptor, generating "EXO-DEPT" EVs. EXO-DEPT treatment specifically enabled HER2-overexpressing BT474 cells to convert CNOB into MCHB in actinomycin D-independent manner, showing successful and specific delivery of HChrR6 mRNA. EXO-DEPTs-but not undirected EVs-plus CNOB caused near-complete growth arrest of orthotopic BT474 xenografts in vivo, demonstrating for the first time EV-mediated delivery of functional exogenous mRNA to tumors. EXO-DEPTs may be generated from patients' own dendritic cells to evade immune rejection, and without plasmids and their potentially harmful genetic material, raising the prospect of clinical use of this regimen. This approach can be used to treat any disease overexpressing a specific marker. Mol Cancer Ther; 17(5); 1133-42. ©2018 AACR.
©2018 American Association for Cancer Research.
Conflict of interest statement
Figures
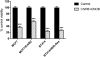

References
-
- Williams EM, Little RF, Mowday AM, et al. Nitroreductase gene-directed enzyme prodrug therapy: insights and advances toward clinical utility. Biochem J. 2015;471:131–153. - PubMed
-
- Barak Y, Thorne SH, Ackerley DF, et al. New enzyme for reductive cancer chemotherapy, YieF, and its improvement by directed evolution. Mol Cancer Ther. 2006;5:97–103. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

