Clonally diverse CD38+HLA-DR+CD8+ T cells persist during fatal H7N9 disease
- PMID: 29483513
- PMCID: PMC5827521
- DOI: 10.1038/s41467-018-03243-7
Clonally diverse CD38+HLA-DR+CD8+ T cells persist during fatal H7N9 disease
Abstract
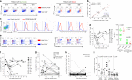
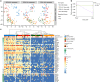
Severe influenza A virus (IAV) infection is associated with immune dysfunction. Here, we show circulating CD8+ T-cell profiles from patients hospitalized with avian H7N9, seasonal IAV, and influenza vaccinees. Patient survival reflects an early, transient prevalence of highly activated CD38+HLA-DR+PD-1+ CD8+ T cells, whereas the prolonged persistence of this set is found in ultimately fatal cases. Single-cell T cell receptor (TCR)-αβ analyses of activated CD38+HLA-DR+CD8+ T cells show similar TCRαβ diversity but differential clonal expansion kinetics in surviving and fatal H7N9 patients. Delayed clonal expansion associated with an early dichotomy at a transcriptome level (as detected by single-cell RNAseq) is found in CD38+HLA-DR+CD8+ T cells from patients who succumbed to the disease, suggesting a divergent differentiation pathway of CD38+HLA-DR+CD8+ T cells from the outset during fatal disease. Our study proposes that effective expansion of cross-reactive influenza-specific TCRαβ clonotypes with appropriate transcriptome signatures is needed for early protection against severe influenza disease.
Conflict of interest statement
The authors declare no competing financial interests.
Figures




References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

