Intestine farnesoid X receptor agonist and the gut microbiota activate G-protein bile acid receptor-1 signaling to improve metabolism
- PMID: 29486523
- PMCID: PMC6111007
- DOI: 10.1002/hep.29857
Intestine farnesoid X receptor agonist and the gut microbiota activate G-protein bile acid receptor-1 signaling to improve metabolism
Abstract
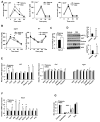
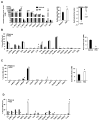
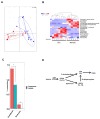
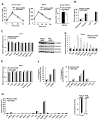
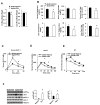
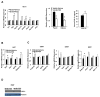
Bile acids activate farnesoid X receptor (FXR) and G protein-coupled bile acid receptor-1 (aka Takeda G protein-coupled receptor-5 [TGR5]) to regulate bile acid metabolism and glucose and insulin sensitivity. FXR and TGR5 are coexpressed in the enteroendocrine L cells, but their roles in integrated regulation of metabolism are not completely understood. We reported recently that activation of FXR induces TGR5 to stimulate glucagon-like peptide-1 (GLP-1) secretion to improve insulin sensitivity and hepatic metabolism. In this study, we used the intestine-restricted FXR agonist fexaramine (FEX) to study the effect of activation of intestinal FXR on the gut microbiome, bile acid metabolism, and FXR and TGR5 signaling. The current study revealed that FEX markedly increased taurolithocholic acid, increased secretion of fibroblast growth factors 15 and 21 and GLP-1, improved insulin and glucose tolerance, and promoted white adipose tissue browning in mice. Analysis of 16S ribosomal RNA sequences of the gut microbiome identified the FEX-induced and lithocholic acid-producing bacteria Acetatifactor and Bacteroides. Antibiotic treatment completely reversed the FEX-induced metabolic phenotypes and inhibited taurolithocholic acid synthesis, adipose tissue browning, and liver bile acid synthesis gene expression but further increased intestinal FXR target gene expression. FEX treatment effectively improved lipid profiles, increased GLP-1 secretion, improved glucose and insulin tolerance, and promoted adipose tissue browning, while antibiotic treatment reversed the beneficial metabolic effects of FEX in obese and diabetic mice.
Conclusion: This study uncovered a mechanism in which activation of intestinal FXR shaped the gut microbiota to activate TGR5/GLP-1 signaling to improve hepatic glucose and insulin sensitivity and increase adipose tissue browning; the gut microbiota plays a critical role in bile acid metabolism and signaling to regulate metabolic homeostasis in health and disease. (Hepatology 2018).
© 2018 by the American Association for the Study of Liver Diseases.
Conflict of interest statement
Figures
Comment in
-
Bile Acids, Microbiota, and Metabolism.Hepatology. 2018 Oct;68(4):1229-1231. doi: 10.1002/hep.30078. Hepatology. 2018. PMID: 29729182 Free PMC article. No abstract available.
References
-
- Lefebvre P, Cariou B, Lien F, Kuipers F, Staels B. Role of bile acids and bile Acid receptors in metabolic regulation. Physiol Rev. 2009;89:147–191. - PubMed
-
- Inagaki T, Choi M, Moschetta A, Peng L, Cummins CL, McDonald JG, Luo G, et al. Fibroblast growth factor 15 functions as an enterohepatic signal to regulate bile acid homeostasis. Cell Metab. 2005;2:217–225. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

