Long Noncoding RNA uc002yug.2 Activates HIV-1 Latency through Regulation of mRNA Levels of Various RUNX1 Isoforms and Increased Tat Expression
- PMID: 29491162
- PMCID: PMC5899182
- DOI: 10.1128/JVI.01844-17
Long Noncoding RNA uc002yug.2 Activates HIV-1 Latency through Regulation of mRNA Levels of Various RUNX1 Isoforms and Increased Tat Expression
Abstract
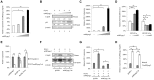
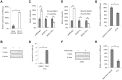
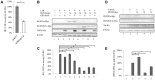
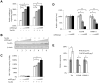
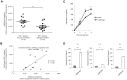
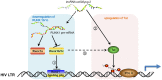
The HIV-1 reservoir is a major obstacle to complete eradication of the virus. Although many proteins and RNAs have been characterized as regulators in HIV-1/AIDS pathogenesis and latency, only a few long noncoding RNAs (lncRNAs) have been shown to be closely associated with HIV-1 replication and latency. In this study, we demonstrated that lncRNA uc002yug.2 plays a key role in HIV-1 replication and latency. uc002yug.2 potentially enhances HIV-1 replication, long terminal repeat (LTR) activity, and the activation of latent HIV-1 in both cell lines and CD4+ T cells from patients. Further investigation revealed that uc002yug.2 activates latent HIV-1 through downregulating RUNX1b and -1c and upregulating Tat protein expression. The accumulated evidence supports our model that the Tat protein has the key role in the uc002yug.2-mediated regulatory effect on HIV-1 reactivation. Moreover, uc002yug.2 showed an ability to activate HIV-1 similar to that of suberoylanilide hydroxamic acid or phorbol 12-myristate 13-acetate using latently infected cell models. These findings improve our understanding of lncRNA regulation of HIV-1 replication and latency, providing new insights into potential targeted therapeutic interventions.IMPORTANCE The latent viral reservoir is the primary obstacle to curing HIV-1 disease. To date, only a few lncRNAs, which play major roles in various biological processes, including viral infection, have been identified as regulators in HIV-1 latency. In this study, we demonstrated that lncRNA uc002yug.2 is important for both HIV-1 replication and activation of latent viruses. Moreover, uc002yug.2 was shown to activate latent HIV-1 through regulating alternative splicing of RUNX1 and increasing the expression of Tat protein. These findings highlight the potential merit of targeting lncRNA uc002yug.2 as an activating agent for latent HIV-1.
Keywords: HIV-1 latency; RUNX1; Tat; activation; lncRNA uc002yug.2.
Copyright © 2018 American Society for Microbiology.
Figures



Similar articles
-
Long noncoding RNA NRON contributes to HIV-1 latency by specifically inducing tat protein degradation.Nat Commun. 2016 Jun 13;7:11730. doi: 10.1038/ncomms11730. Nat Commun. 2016. PMID: 27291871 Free PMC article.
-
NF-κB-Interacting Long Noncoding RNA Regulates HIV-1 Replication and Latency by Repressing NF-κB Signaling.J Virol. 2020 Aug 17;94(17):e01057-20. doi: 10.1128/JVI.01057-20. Print 2020 Aug 17. J Virol. 2020. PMID: 32581100 Free PMC article.
-
Posttranscriptional Regulation of HIV-1 Gene Expression during Replication and Reactivation from Latency by Nuclear Matrix Protein MATR3.mBio. 2018 Nov 13;9(6):e02158-18. doi: 10.1128/mBio.02158-18. mBio. 2018. PMID: 30425153 Free PMC article.
-
HIV Tat as a latency reversing agent: turning the tables on viral persistence.Front Immunol. 2025 Apr 11;16:1571151. doi: 10.3389/fimmu.2025.1571151. eCollection 2025. Front Immunol. 2025. PMID: 40292298 Free PMC article. Review.
-
The Molecular Basis for Human Immunodeficiency Virus Latency.Annu Rev Virol. 2017 Sep 29;4(1):261-285. doi: 10.1146/annurev-virology-101416-041646. Epub 2017 Jul 17. Annu Rev Virol. 2017. PMID: 28715973 Review.
Cited by
-
Non-Coding RNAs in HIV Infection, NeuroHIV, and Related Comorbidities.Cells. 2024 May 23;13(11):898. doi: 10.3390/cells13110898. Cells. 2024. PMID: 38891030 Free PMC article. Review.
-
A Novel lncRNA, AK130181, Contributes to HIV-1 Latency by Regulating Viral Promoter-Driven Gene Expression in Primary CD4+ T Cells.Mol Ther Nucleic Acids. 2020 Jun 5;20:754-763. doi: 10.1016/j.omtn.2020.04.011. Epub 2020 Apr 29. Mol Ther Nucleic Acids. 2020. PMID: 32408053 Free PMC article.
-
Human splice factors contribute to latent HIV infection in primary cell models and blood CD4+ T cells from ART-treated individuals.PLoS Pathog. 2020 Nov 30;16(11):e1009060. doi: 10.1371/journal.ppat.1009060. eCollection 2020 Nov. PLoS Pathog. 2020. PMID: 33253324 Free PMC article.
-
Roles of lncRNAs in the transcription regulation of HIV-1.Biomed J. 2022 Aug;45(4):580-593. doi: 10.1016/j.bj.2022.03.012. Epub 2022 Mar 29. Biomed J. 2022. PMID: 35364293 Free PMC article. Review.
-
Non-Coding RNAs in the Etiology and Control of Major and Neglected Human Tropical Diseases.Front Immunol. 2021 Oct 19;12:703936. doi: 10.3389/fimmu.2021.703936. eCollection 2021. Front Immunol. 2021. PMID: 34737736 Free PMC article. Review.
References
-
- Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, Cabili MN, Jaenisch R, Mikkelsen TS, Jacks T, Hacohen N, Bernstein BE, Kellis M, Regev A, Rinn JL, Lander ES. 2009. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458:223–227. doi:10.1038/nature07672. - DOI - PMC - PubMed
-
- Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D, Thomas K, Presser A, Bernstein BE, van Oudenaarden A, Regev A, Lander ES, Rinn JL. 2009. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci U S A 106:11667–11672. doi:10.1073/pnas.0904715106. - DOI - PMC - PubMed
-
- Gutschner T, Hammerle M, Eissmann M, Hsu J, Kim Y, Hung G, Revenko A, Arun G, Stentrup M, Gross M, Zornig M, MacLeod AR, Spector DL, Diederichs S. 2013. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res 73:1180–1189. doi:10.1158/0008-5472.CAN-12-2850. - DOI - PMC - PubMed
-
- Han Y, Liu Y, Nie L, Gui Y, Cai Z. 2013. Inducing cell proliferation inhibition, apoptosis, and motility reduction by silencing long noncoding ribonucleic acid metastasis-associated lung adenocarcinoma transcript 1 in urothelial carcinoma of the bladder. Urology 81:209.e1–209.e7. doi:10.1016/j.urology.2012.08.044. - DOI - PubMed
-
- Pandey GK, Mitra S, Subhash S, Hertwig F, Kanduri M, Mishra K, Fransson S, Ganeshram A, Mondal T, Bandaru S, Ostensson M, Akyurek LM, Abrahamsson J, Pfeifer S, Larsson E, Shi L, Peng Z, Fischer M, Martinsson T, Hedborg F, Kogner P, Kanduri C. 2014. The risk-associated long noncoding RNA NBAT-1 controls neuroblastoma progression by regulating cell proliferation and neuronal differentiation. Cancer Cell 26:722–737. doi:10.1016/j.ccell.2014.09.014. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

