A multiplatform strategy for the discovery of conventional monoclonal antibodies that inhibit the voltage-gated potassium channel Kv1.3
- PMID: 29494279
- PMCID: PMC5973702
- DOI: 10.1080/19420862.2018.1445451
A multiplatform strategy for the discovery of conventional monoclonal antibodies that inhibit the voltage-gated potassium channel Kv1.3
Abstract
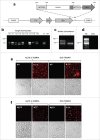
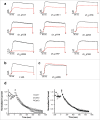
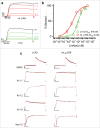
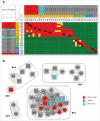
Identifying monoclonal antibodies that block human voltage-gated ion channels (VGICs) is a challenging endeavor exacerbated by difficulties in producing recombinant ion channel proteins in amounts that support drug discovery programs. We have developed a general strategy to address this challenge by combining high-level expression of recombinant VGICs in Tetrahymena thermophila with immunization of phylogenetically diverse species and unique screening tools that allow deep-mining for antibodies that could potentially bind functionally important regions of the protein. Using this approach, we targeted human Kv1.3, a voltage-gated potassium channel widely recognized as a therapeutic target for the treatment of a variety of T-cell mediated autoimmune diseases. Recombinant Kv1.3 was used to generate and recover 69 full-length anti-Kv1.3 mAbs from immunized chickens and llamas, of which 10 were able to inhibit Kv1.3 current. Select antibodies were shown to be potent (IC50<10 nM) and specific for Kv1.3 over related Kv1 family members, hERG and hNav1.5.
Keywords: Kv1.3; Tetrahymena; antibody; voltage-gated ion channel.
Figures


References
-
- Onimura H, Ohki K, Nozawa Y, Naitoh Y. Electrical properties of Tetrahymena, a suitable tool for studies on membrane excitation. Proc. Japan. Acad. 1980;56:538–43. doi:10.2183/pjab.56.538. - DOI
-
- Beeton C, Wulff H, Standifer NE, Azam P, Mullen KM, Pennington MW, Kolski-Andreaco A, Wei E, Grino A, Counts DR, et al.. Kv1.3 channels are a therapeutic target for T cell-mediated autoimmune diseases. Proc Natl Acad Sci USA. 2006;103:17414–9. doi:10.1073/pnas.0605136103. PMID:17088564. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
