Amino acid-level signal-to-noise analysis of incidentally identified variants in genes associated with long QT syndrome during pediatric whole exome sequencing reflects background genetic noise
- PMID: 29501670
- PMCID: PMC6026069
- DOI: 10.1016/j.hrthm.2018.02.031
Amino acid-level signal-to-noise analysis of incidentally identified variants in genes associated with long QT syndrome during pediatric whole exome sequencing reflects background genetic noise
Abstract
Background: Due to rapid expansion of clinical genetic testing, an increasing number of genetic variants of undetermined significance and unclear diagnostic value are being identified in children. Variants found in genes associated with heritable channelopathies, such as long QT syndrome (LQTS), are particularly difficult to interpret given the risk of sudden cardiac death associated with pathologic mutations.
Objective: The purpose of this study was to determine whether variants in LQTS-associated genes from whole exome sequencing (WES) represent disease-associated biomarkers or background genetic "noise."
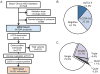
Methods: WES variants from Baylor Genetics Laboratories were obtained for 17 LQTS-associated genes. Rare variants from healthy controls were obtained from the GnomAD database. LQTS case variants were extracted from the literature. Amino acid-level mapping and signal-to-noise calculations were conducted. Clinical history and diagnostic studies were analyzed for WES subjects evaluated at our institution.
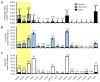
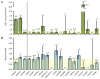
Results: Variants in LQTS case-associated genes were present in 38.3% of 7244 WES probands. There was a similar frequency of variants in the WES and healthy cohorts for LQTS1-3 (11.2% and 12.9%, respectively) and LQTS4-17 (27.1% and 38.4%, respectively). WES variants preferentially localized to amino acids altered in control individuals compared to cases. Based on amino acid-level analysis, WES-identified variants are indistinguishable from healthy background variation, whereas LQTS1 and 2 case-identified variants localized to clear pathologic "hotspots." No individuals who underwent clinical evaluation had clinical suspicion for LQTS.
Conclusion: The prevalence of incidentally identified LQTS-associated variants is ∼38% among WES tests. These variants most likely represent benign healthy background genetic variation rather than disease-associated mutations.
Keywords: Genetic testing; Genetics; Long QT syndrome; Mutation; Variant of undetermined significance; Whole exome sequencing.
Copyright © 2018 Heart Rhythm Society. Published by Elsevier Inc. All rights reserved.
Figures
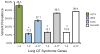

Similar articles
-
Incidentally identified genetic variants in arrhythmogenic right ventricular cardiomyopathy-associated genes among children undergoing exome sequencing reflect healthy population variation.Mol Genet Genomic Med. 2019 Jun;7(6):e593. doi: 10.1002/mgg3.593. Epub 2019 Apr 15. Mol Genet Genomic Med. 2019. PMID: 30985088 Free PMC article.
-
Interpreting Incidentally Identified Variants in Genes Associated With Catecholaminergic Polymorphic Ventricular Tachycardia in a Large Cohort of Clinical Whole-Exome Genetic Test Referrals.Circ Arrhythm Electrophysiol. 2017 Apr;10(4):e004742. doi: 10.1161/CIRCEP.116.004742. Circ Arrhythm Electrophysiol. 2017. PMID: 28404607 Free PMC article.
-
Semiconductor Whole Exome Sequencing for the Identification of Genetic Variants in Colombian Patients Clinically Diagnosed with Long QT Syndrome.Mol Diagn Ther. 2016 Aug;20(4):353-62. doi: 10.1007/s40291-016-0207-2. Mol Diagn Ther. 2016. PMID: 27251404
-
Genetics of long QT syndrome.Methodist Debakey Cardiovasc J. 2014 Jan-Mar;10(1):29-33. doi: 10.14797/mdcj-10-1-29. Methodist Debakey Cardiovasc J. 2014. PMID: 24932360 Free PMC article. Review.
-
Modern methods in diagnostics and research of molecular bases of rare diseases.Cas Lek Cesk. 2018 Summer;157(3):133-136. Cas Lek Cesk. 2018. PMID: 30441942 Review. English.
Cited by
-
Analysis of enriched rare variants in JPH2-encoded junctophilin-2 among Greater Middle Eastern individuals reveals a novel homozygous variant associated with neonatal dilated cardiomyopathy.Sci Rep. 2019 Jun 21;9(1):9038. doi: 10.1038/s41598-019-44987-6. Sci Rep. 2019. PMID: 31227780 Free PMC article.
-
DiscoVari: A Web-Based Precision Medicine Tool for Predicting Variant Pathogenicity in Cardiomyopathy- and Channelopathy-Associated Genes.Circ Genom Precis Med. 2023 Aug;16(4):317-327. doi: 10.1161/CIRCGEN.122.003911. Epub 2023 Jul 6. Circ Genom Precis Med. 2023. PMID: 37409478 Free PMC article.
-
Incidentally identified genetic variants in arrhythmogenic right ventricular cardiomyopathy-associated genes among children undergoing exome sequencing reflect healthy population variation.Mol Genet Genomic Med. 2019 Jun;7(6):e593. doi: 10.1002/mgg3.593. Epub 2019 Apr 15. Mol Genet Genomic Med. 2019. PMID: 30985088 Free PMC article.
-
GENESIS: Gene-Specific Machine Learning Models for Variants of Uncertain Significance Found in Catecholaminergic Polymorphic Ventricular Tachycardia and Long QT Syndrome-Associated Genes.Circ Arrhythm Electrophysiol. 2022 Apr;15(4):e010326. doi: 10.1161/CIRCEP.121.010326. Epub 2022 Mar 31. Circ Arrhythm Electrophysiol. 2022. PMID: 35357185 Free PMC article.
-
From diagnostic testing to precision medicine: the evolving role of genomics in cardiac channelopathies and cardiomyopathies in children.Curr Opin Genet Dev. 2022 Oct;76:101978. doi: 10.1016/j.gde.2022.101978. Epub 2022 Sep 1. Curr Opin Genet Dev. 2022. PMID: 36058060 Free PMC article. Review.
References
-
- Landstrom AP, Tester DJ, Ackerman MJ. Role of genetic testing for sudden death predisposing heart conditions in athletes. 2011
-
- Wang Q, Curran ME, Splawski I, et al. Positional cloning of a novel potassium channel gene: KVLQT1 mutations cause cardiac arrhythmias. Nat Genet. 1996;12:17–23. - PubMed
-
- Curran ME, Splawski I, Timothy KW, Vincent GM, Green ED, Keating MT. A molecular basis for cardiac arrhythmia: HERG mutations cause long QT syndrome. Cell. 1995;80:795–803. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

