Design, synthesis, and biological evaluation of stable β6.3-Helices: Discovery of non-hemolytic antibacterial peptides
- PMID: 29501941
- PMCID: PMC8366898
- DOI: 10.1016/j.ejmech.2018.02.057
Design, synthesis, and biological evaluation of stable β6.3-Helices: Discovery of non-hemolytic antibacterial peptides
Abstract
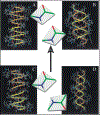
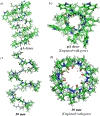
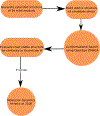
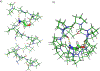
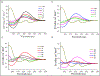
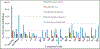
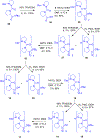
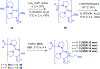
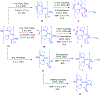
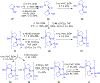
Gramicidin A, a topical antibiotic made from alternating L and D amino acids, is characterized by its wide central pore; upon insertion into membranes, it forms channels that disrupts ion gradients. We present helical peptidomimetics with this characteristic wide central pore that have been designed to mimic gramicidin A channels. Mimetics were designed using molecular modeling focused on oligomers of heterochiral dipeptides of proline analogs, in particular azaproline (AzPro). Molecular Dynamics simulations in water confirmed the stability of the designed helices. A sixteen-residue Formyl-(AzPro-Pro)8-NHCH2CH2OH helix was synthesized as well as a full thirty-two residue Cbz-(AzPro-Pro)16-OtBu channels. No liposomal lysis activity was observed suggesting lack of channel formation, possibly due to inappropriate hydrogen-bonding interactions in the membrane. These peptidomimetics also did not hemolyze red blood cells, unlike gramicidin A.
Keywords: Antibiotic; Azaproline foldamers; Gramicidin; Liposomes and beta-helices; RBC hemolysis.
Copyright © 2018 Elsevier Masson SAS. All rights reserved.
Figures
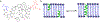




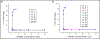


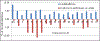

Similar articles
-
Gramicidin-S-Inspired Cyclopeptidomimetics as Potent Membrane-Active Bactericidal Agents with Therapeutic Potential.ChemMedChem. 2021 Jan 19;16(2):368-376. doi: 10.1002/cmdc.202000568. Epub 2020 Oct 16. ChemMedChem. 2021. PMID: 33026182
-
Antibacterial evaluation of structurally amphipathic, membrane active small cationic peptidomimetics: synthesized by incorporating 3-amino benzoic acid as peptidomimetic element.Eur J Med Chem. 2014 Aug 18;83:102-15. doi: 10.1016/j.ejmech.2014.06.023. Epub 2014 Jun 12. Eur J Med Chem. 2014. PMID: 24953028
-
Rational design, synthesis, and biological evaluation of lactam-bridged gramicidin A analogues: discovery of a low-hemolytic antibacterial peptide.ChemMedChem. 2015 Mar;10(3):540-5. doi: 10.1002/cmdc.201402473. Epub 2014 Dec 15. ChemMedChem. 2015. PMID: 25510221
-
Limiting Assumptions in the Design of Peptidomimetics.Drug Dev Res. 2017 Sep;78(6):245-267. doi: 10.1002/ddr.21406. Epub 2017 Sep 6. Drug Dev Res. 2017. PMID: 28875546 Review.
-
Design, synthesis and evaluation of bioactivity of peptidomimetics based on chloroalkene dipeptide isosteres.Bioorg Med Chem. 2024 Aug 1;110:117811. doi: 10.1016/j.bmc.2024.117811. Epub 2024 Jun 24. Bioorg Med Chem. 2024. PMID: 38959684 Review.
Cited by
-
Fluorine-hydrogen inter-actions observed in a helix structure having an orn-free gramicidin S sequence incorporating 4-trans-fluoro-proline.Acta Crystallogr E Crystallogr Commun. 2025 Mar 27;81(Pt 4):345-349. doi: 10.1107/S2056989025002592. eCollection 2025 Apr 1. Acta Crystallogr E Crystallogr Commun. 2025. PMID: 40201012 Free PMC article.
-
Bio-evaluation of fluoro and trifluoromethyl-substituted salicylanilides against multidrug-resistant S. aureus.Med Chem Res. 2021;30(12):2301-2315. doi: 10.1007/s00044-021-02808-4. Epub 2021 Oct 27. Med Chem Res. 2021. PMID: 34720564 Free PMC article.
References
-
- Cabezas E, Satterthwait AC, The hydrogen bond mimic approach: solid-phase synthesis of a peptide stabilized as an α-helix with a hydrazone link, J. Am. Chem. Soc, 121 (1999) 3862–3875.
-
- Chapman RN, Dimartino G, Arora PS, A highly stable short α-helix constrained by a main-chain hydrogen-bond surrogate, J. Am. Chem. Soc, 126 (2004) 12252–12253. - PubMed
-
- Stanger HE, Gellman SH, Rules for antiparallel β-sheet design: D-Pro-Gly is superior to L-Asn-Gly for β-hairpin Nucleation, J. Am. Chem. Soc, 120 (1998) 4236–4237.
-
- Schneider JP, Kelly JW, Templates that induce α-helical, β-sheet, and loop conformations, Chem. Rev, 95 (1995) 2169–2187.
-
- Kemp DS, Bowen BR, Muendel CC, Synthesis and conformational analysis of epindolidione-derived peptide models for β-sheet formation, J. Org. Chem, 55 (1990) 4650–4657.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

