Efficient Non-viral Gene Delivery into Human Hematopoietic Stem Cells by Minicircle Sleeping Beauty Transposon Vectors
- PMID: 29503198
- PMCID: PMC6079369
- DOI: 10.1016/j.ymthe.2018.01.012
Efficient Non-viral Gene Delivery into Human Hematopoietic Stem Cells by Minicircle Sleeping Beauty Transposon Vectors
Abstract
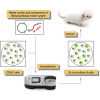
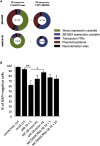
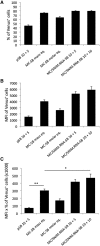
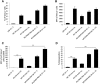
The Sleeping Beauty (SB) transposon system is a non-viral gene delivery platform that combines simplicity, inexpensive manufacture, and favorable safety features in the context of human applications. However, efficient correction of hematopoietic stem and progenitor cells (HSPCs) with non-viral vector systems, including SB, demands further refinement of gene delivery techniques. We set out to improve SB gene transfer into hard-to-transfect human CD34+ cells by vectorizing the SB system components in the form of minicircles that are devoid of plasmid backbone sequences and are, therefore, significantly reduced in size. As compared to conventional plasmids, delivery of the SB transposon system as minicircle DNA is ∼20 times more efficient, and it is associated with up to a 50% reduction in cellular toxicity in human CD34+ cells. Moreover, providing the SB transposase in the form of synthetic mRNA enabled us to further increase the efficacy and biosafety of stable gene delivery into hematopoietic progenitors ex vivo. Genome-wide insertion site profiling revealed a close-to-random distribution of SB transposon integrants, which is characteristically different from gammaretroviral and lentiviral integrations in HSPCs. Transplantation of gene-marked CD34+ cells in immunodeficient mice resulted in long-term engraftment and hematopoietic reconstitution, which was most efficient when the SB transposase was supplied as mRNA and nucleofected cells were maintained for 4-8 days in culture before transplantation. Collectively, implementation of minicircle and mRNA technologies allowed us to further refine the SB transposon system in the context of HSPC gene delivery to ultimately meet clinical demands of an efficient and safe non-viral gene therapy protocol.
Keywords: chromosomal integration; gene therapy; gene vectors; hematopoietic stem cells; nonviral gene delivery; transposition.
Copyright © 2018 The Authors. Published by Elsevier Inc. All rights reserved.
Figures
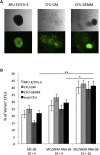
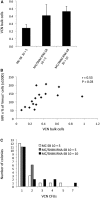
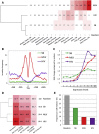
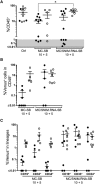
Similar articles
-
Stable transgene expression in primitive human CD34+ hematopoietic stem/progenitor cells, using the Sleeping Beauty transposon system.Hum Gene Ther. 2009 Dec;20(12):1607-26. doi: 10.1089/hum.2009.109. Hum Gene Ther. 2009. PMID: 19689196 Free PMC article.
-
Efficient stable gene transfer into human cells by the Sleeping Beauty transposon vectors.Methods. 2009 Nov;49(3):287-97. doi: 10.1016/j.ymeth.2009.07.001. Epub 2009 Jul 15. Methods. 2009. PMID: 19615447 Review.
-
Transposons: Moving Forward from Preclinical Studies to Clinical Trials.Hum Gene Ther. 2017 Nov;28(11):1087-1104. doi: 10.1089/hum.2017.128. Epub 2017 Aug 22. Hum Gene Ther. 2017. PMID: 28920716 Review.
-
Sleeping Beauty transposition: from biology to applications.Crit Rev Biochem Mol Biol. 2017 Feb;52(1):18-44. doi: 10.1080/10409238.2016.1237935. Epub 2016 Oct 4. Crit Rev Biochem Mol Biol. 2017. PMID: 27696897 Review.
-
Gene transfer into genomes of human cells by the sleeping beauty transposon system.Mol Ther. 2003 Jul;8(1):108-17. doi: 10.1016/s1525-0016(03)00099-6. Mol Ther. 2003. PMID: 12842434
Cited by
-
Engineered Sleeping Beauty transposase redirects transposon integration away from genes.Nucleic Acids Res. 2022 Mar 21;50(5):2807-2825. doi: 10.1093/nar/gkac092. Nucleic Acids Res. 2022. PMID: 35188569 Free PMC article.
-
At the Intersection of Biomaterials and Gene Therapy: Progress in Non-viral Delivery of Nucleic Acids.Front Bioeng Biotechnol. 2019 Jun 4;7:131. doi: 10.3389/fbioe.2019.00131. eCollection 2019. Front Bioeng Biotechnol. 2019. PMID: 31214586 Free PMC article. Review.
-
Current Non-Viral-Based Strategies to Manufacture CAR-T Cells.Int J Mol Sci. 2024 Dec 21;25(24):13685. doi: 10.3390/ijms252413685. Int J Mol Sci. 2024. PMID: 39769449 Free PMC article. Review.
-
Improving therapeutic potential of non-viral minimized DNA vectors.Cell Gene Ther Insights. 2020 Nov;6(10):1489-1505. doi: 10.18609/cgti.2020.163. Epub 2020 Nov 19. Cell Gene Ther Insights. 2020. PMID: 33953961 Free PMC article.
-
Targeted Transposition of Minicircle DNA Using Single-Chain Antibody Conjugated Cyclodextrin-Modified Poly (Propylene Imine) Nanocarriers.Cancers (Basel). 2022 Apr 11;14(8):1925. doi: 10.3390/cancers14081925. Cancers (Basel). 2022. PMID: 35454835 Free PMC article.
References
-
- Booth C., Gaspar H.B., Thrasher A.J. Treating Immunodeficiency through HSC Gene Therapy. Trends Mol. Med. 2016;22:317–327. - PubMed
-
- Hacein-Bey-Abina S., Von Kalle C., Schmidt M., McCormack M.P., Wulffraat N., Leboulch P., Lim A., Osborne C.S., Pawliuk R., Morillon E. LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1. Science. 2003;302:415–419. - PubMed
-
- Ott M.G., Schmidt M., Schwarzwaelder K., Stein S., Siler U., Koehl U., Glimm H., Kühlcke K., Schilz A., Kunkel H. Correction of X-linked chronic granulomatous disease by gene therapy, augmented by insertional activation of MDS1-EVI1, PRDM16 or SETBP1. Nat. Med. 2006;12:401–409. - PubMed
-
- Howe S.J., Mansour M.R., Schwarzwaelder K., Bartholomae C., Hubank M., Kempski H., Brugman M.H., Pike-Overzet K., Chatters S.J., de Ridder D. Insertional mutagenesis combined with acquired somatic mutations causes leukemogenesis following gene therapy of SCID-X1 patients. J. Clin. Invest. 2008;118:3143–3150. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

