Impact of Phosphoproteomics in the Era of Precision Medicine for Prostate Cancer
- PMID: 29503809
- PMCID: PMC5820335
- DOI: 10.3389/fonc.2018.00028
Impact of Phosphoproteomics in the Era of Precision Medicine for Prostate Cancer
Abstract
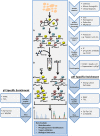
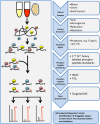
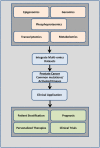
Prostate cancer is the most common malignancy in men in the United States. While androgen deprivation therapy results in tumor responses initially, there is relapse and progression to metastatic castration-resistant prostate cancer. Currently, all prostate cancer patients receive essentially the same treatment, and there is a need for clinically applicable technologies to provide predictive biomarkers toward personalized therapies. Genomic analyses of tumors are used for clinical applications, but with a paucity of obvious driver mutations in metastatic castration-resistant prostate cancer, other applications, such as phosphoproteomics, may complement this approach. Immunohistochemistry and reverse phase protein arrays are limited by the availability of reliable antibodies and evaluates a preselected number of targets. Mass spectrometry-based phosphoproteomics has been used to profile tumors consisting of thousands of phosphopeptides from individual patients after surgical resection or at autopsy. However, this approach is time consuming, and while a large number of candidate phosphopeptides are obtained for evaluation, limitations are reduced reproducibility, sensitivity, and precision. Targeted mass spectrometry can help eliminate these limitations and is more cost effective and less time consuming making it a practical platform for future clinical testing. In this review, we discuss the use of phosphoproteomics in prostate cancer and other clinical cancer tissues for target identification, hypothesis testing, and possible patient stratification. We highlight the majority of studies that have used phosphoproteomics in prostate cancer tissues and cell lines and propose ways forward to apply this approach in basic and clinical research. Overall, the implementation of phosphoproteomics via targeted mass spectrometry has tremendous potential to aid in the development of more rational, personalized therapies that will result in increased survival and quality of life enhancement in patients suffering from metastatic castration-resistant prostate cancer.
Keywords: clinical trials; kinase inhibitors; kinases; mass spectrometry; phosphoproteomics; prostate cancer; signaling pathways; targeted mass spectrometry.
Figures


Similar articles
-
Personalization of prostate cancer therapy through phosphoproteomics.Nat Rev Urol. 2018 Aug;15(8):483-497. doi: 10.1038/s41585-018-0014-0. Nat Rev Urol. 2018. PMID: 29752463 Review.
-
Phosphotyrosine-based-phosphoproteomics scaled-down to biopsy level for analysis of individual tumor biology and treatment selection.J Proteomics. 2017 Jun 6;162:99-107. doi: 10.1016/j.jprot.2017.04.014. Epub 2017 Apr 23. J Proteomics. 2017. PMID: 28442448
-
Identifying therapeutic strategies for triple-negative breast cancer via phosphoproteomics.Expert Rev Proteomics. 2024 Nov 26:1-17. doi: 10.1080/14789450.2024.2432477. Online ahead of print. Expert Rev Proteomics. 2024. PMID: 39588933 Review.
-
Integrating phosphoproteomics into kinase-targeted cancer therapies in precision medicine.J Proteomics. 2019 Jan 16;191:68-79. doi: 10.1016/j.jprot.2018.03.033. Epub 2018 Apr 3. J Proteomics. 2019. PMID: 29621648 Review.
-
Update on Systemic Prostate Cancer Therapies: Management of Metastatic Castration-resistant Prostate Cancer in the Era of Precision Oncology.Eur Urol. 2019 Jan;75(1):88-99. doi: 10.1016/j.eururo.2018.03.028. Epub 2018 Apr 16. Eur Urol. 2019. PMID: 29673712 Review.
Cited by
-
Ion Activation Methods for Peptides and Proteins.Anal Chem. 2020 Jan 7;92(1):227-251. doi: 10.1021/acs.analchem.9b04859. Epub 2019 Nov 12. Anal Chem. 2020. PMID: 31665881 Free PMC article. Review. No abstract available.
-
Optical chemosensors for the detection of proximally phosphorylated peptides and proteins.RSC Chem Biol. 2021 Apr 21;2(3):815-829. doi: 10.1039/d1cb00055a. eCollection 2021 Jun 1. RSC Chem Biol. 2021. PMID: 34458812 Free PMC article. Review.
-
Strategies for Functional Interrogation of Big Cancer Data Using Drosophila Cancer Models.Int J Mol Sci. 2020 May 26;21(11):3754. doi: 10.3390/ijms21113754. Int J Mol Sci. 2020. PMID: 32466549 Free PMC article. Review.
-
Proteomic Landscape of Prostate Cancer: The View Provided by Quantitative Proteomics, Integrative Analyses, and Protein Interactomes.Cancers (Basel). 2021 Sep 27;13(19):4829. doi: 10.3390/cancers13194829. Cancers (Basel). 2021. PMID: 34638309 Free PMC article. Review.
-
Halting the vicious cycle within the multiple myeloma ecosystem: blocking JAM-A on bone marrow endothelial cells restores angiogenic homeostasis and suppresses tumor progression.Haematologica. 2021 Jul 1;106(7):1943-1956. doi: 10.3324/haematol.2019.239913. Haematologica. 2021. PMID: 32354870 Free PMC article.
References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

