Life times of metastable states guide regulatory signaling in transcriptional riboswitches
- PMID: 29507289
- PMCID: PMC5838219
- DOI: 10.1038/s41467-018-03375-w
Life times of metastable states guide regulatory signaling in transcriptional riboswitches
Abstract
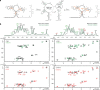
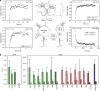
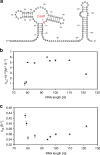
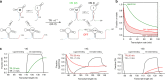
Transcriptional riboswitches modulate downstream gene expression by a tight coupling of ligand-dependent RNA folding kinetics with the rate of transcription. RNA folding pathways leading to functional ON and OFF regulation involve the formation of metastable states within well-defined sequence intervals during transcription. The kinetic requirements for the formation and preservation of these metastable states in the context of transcription remain unresolved. Here, we reversibly trap the previously defined regulatory relevant metastable intermediate of the Mesoplasma florum 2'-deoxyguanosine (2'dG)-sensing riboswitch using a photocaging-ligation approach, and monitor folding to its native state by real-time NMR in both presence and absence of ligand. We further determine transcription rates for two different bacterial RNA polymerases. Our results reveal that the riboswitch functions only at transcription rates typical for bacterial polymerases (10-50 nt s-1) and that gene expression is modulated by 40-50% only, while subtle differences in folding rates guide population ratios within the structural ensemble to a specific regulatory outcome.
Conflict of interest statement
The authors declare no competing interests.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

