A Nogo-Like Signaling Perspective from Birth to Adulthood and in Old Age: Brain Expression Patterns of Ligands, Receptors and Modulators
- PMID: 29520216
- PMCID: PMC5827527
- DOI: 10.3389/fnmol.2018.00042
A Nogo-Like Signaling Perspective from Birth to Adulthood and in Old Age: Brain Expression Patterns of Ligands, Receptors and Modulators
Abstract
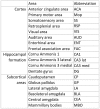
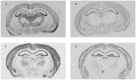
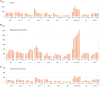
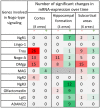
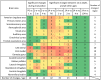
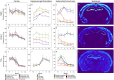
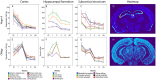
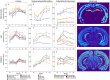
An appropriate strength of Nogo-like signaling is important to maintain synaptic homeostasis in the CNS. Disturbances have been associated with schizophrenia, MS and other diseases. Blocking Nogo-like signaling may improve recovery after spinal cord injury, stroke and traumatic brain injury. To understand the interacting roles of an increasing number of ligands, receptors and modulators engaged in Nogo-like signaling, the transcriptional activity of these genes in the same brain areas from birth to old age in the normal brain is needed. Thus, we have quantitatively mapped the innate expression of 11 important genes engaged in Nogo-like signaling. Using in situ hybridization, we located and measured the amount of mRNA encoding Nogo-A, OMgp, NgR1, NgR2, NgR3, Lingo-1, Troy, Olfactomedin, LgI1, ADAM22, and MAG, in 18 different brain areas at six different ages (P0, 1, 2, 4, 14, and 104 weeks). We show gene- and area-specific activities and how the genes undergo dynamic regulation during postnatal development and become stable during adulthood. Hippocampal areas underwent the largest changes over time. We only found differences between individual cortical areas in Troy and MAG. Subcortical areas presented the largest inter-regional differences; lateral and basolateral amygdala had markedly higher expression than other subcortical areas. The widespread differences and unique expression patterns of the different genes involved in Nogo-like signaling suggest that the functional complexes could look vastly different in different areas.
Keywords: Lingo-1; NgR1; Nogo-A; OMgp; Olfactomedin; Troy; development; hippocampus.
Figures





Similar articles
-
Spatiotemporal and Long Lasting Modulation of 11 Key Nogo Signaling Genes in Response to Strong Neuroexcitation.Front Mol Neurosci. 2017 Apr 11;10:94. doi: 10.3389/fnmol.2017.00094. eCollection 2017. Front Mol Neurosci. 2017. PMID: 28442990 Free PMC article.
-
Differential conserted activity induced regulation of Nogo receptors (1-3), LOTUS and Nogo mRNA in mouse brain.PLoS One. 2013 Apr 11;8(4):e60892. doi: 10.1371/journal.pone.0060892. Print 2013. PLoS One. 2013. PMID: 23593344 Free PMC article.
-
The Nogo-66 receptor homolog NgR2 is a sialic acid-dependent receptor selective for myelin-associated glycoprotein.J Neurosci. 2005 Jan 26;25(4):808-22. doi: 10.1523/JNEUROSCI.4464-04.2005. J Neurosci. 2005. PMID: 15673660 Free PMC article.
-
Targeting the Nogo-A signalling pathway to promote recovery following acute CNS injury.Curr Pharm Des. 2007;13(24):2470-84. doi: 10.2174/138161207781368611. Curr Pharm Des. 2007. PMID: 17692015 Review.
-
The Nogo receptor, its ligands and axonal regeneration in the spinal cord; a review.J Neurocytol. 2002 Feb;31(2):93-120. doi: 10.1023/a:1023941421781. J Neurocytol. 2002. PMID: 12815233 Review.
Cited by
-
Co-Expression of Nogo-A in Dopaminergic Neurons of the Human Substantia Nigra Pars Compacta Is Reduced in Parkinson's Disease.Cells. 2021 Nov 30;10(12):3368. doi: 10.3390/cells10123368. Cells. 2021. PMID: 34943877 Free PMC article.
-
Therapeutic Strategies Targeting Respiratory Recovery after Spinal Cord Injury: From Preclinical Development to Clinical Translation.Cells. 2023 May 31;12(11):1519. doi: 10.3390/cells12111519. Cells. 2023. PMID: 37296640 Free PMC article. Review.
-
Exercise-induced Nogo-A influences rodent motor learning in a time-dependent manner.PLoS One. 2021 May 5;16(5):e0250743. doi: 10.1371/journal.pone.0250743. eCollection 2021. PLoS One. 2021. PMID: 33951058 Free PMC article.
-
The LGI1 protein: molecular structure, physiological functions and disruption-related seizures.Cell Mol Life Sci. 2021 Dec 30;79(1):16. doi: 10.1007/s00018-021-04088-y. Cell Mol Life Sci. 2021. PMID: 34967933 Free PMC article. Review.
-
NgR1 binding to reovirus reveals an unusual bivalent interaction and a new viral attachment protein.Proc Natl Acad Sci U S A. 2023 Jun 13;120(24):e2219404120. doi: 10.1073/pnas.2219404120. Epub 2023 Jun 5. Proc Natl Acad Sci U S A. 2023. PMID: 37276413 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

