Global spatial analysis of Arabidopsis natural variants implicates 5'UTR splicing of LATE ELONGATED HYPOCOTYL in responses to temperature
- PMID: 29520807
- PMCID: PMC6033021
- DOI: 10.1111/pce.13188
Global spatial analysis of Arabidopsis natural variants implicates 5'UTR splicing of LATE ELONGATED HYPOCOTYL in responses to temperature
Abstract
How plants perceive and respond to temperature remains an important question in the plant sciences. Temperature perception and signal transduction may occur through temperature-sensitive intramolecular folding of primary mRNA transcripts. Recent studies suggested a role for retention of the first intron in the 5'UTR of the clock component LATE ELONGATED HYPOCOTYL (LHY) in response to changes in temperature. Here, we identified a set of haplotypes in the LHY 5'UTR, examined their global spatial distribution, and obtained evidence that haplotype can affect temperature-dependent splicing of LHY transcripts. Correlations of haplotype spatial distributions with global bioclimatic variables and altitude point to associations with annual mean temperature and temperature fluctuation. Relatively rare relict type accessions correlate with lower mean temperature and greater temperature fluctuation and the spatial distribution of other haplotypes may be informative of evolutionary processes driving colonization of ecosystems. We propose that haplotypes may possess distinct 5'UTR pre-mRNA folding thermodynamics and/or specific biological stabilities based around the binding of trans-acting RNA splicing factors, a consequence of which is scalable splicing sensitivity of a central clock component that is likely tuned to specific temperature environments.
Keywords: 5′UTR; Arabidopsis; RNA; alternative splicing; circadian clock; natural variation; temperature; thermosensor.
© 2018 The Authors Plant, Cell & Environment Published by John Wiley & Sons Ltd.
Figures



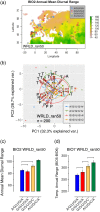


References
-
- Arana, M. V. , Tognacca, R. S. , Estravis‐Barcala, M. , Sanchez, R. A. , & Botto, J. F. (2017). Physiological and molecular mechanisms underlying the integration of light and temperature cues in Arabidopsis thaliana seeds. Plant, Cell & Environment, 40, 3113–3121. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

