Subcellular Organization: A Critical Feature of Bacterial Cell Replication
- PMID: 29522747
- PMCID: PMC5870143
- DOI: 10.1016/j.cell.2018.01.014
Subcellular Organization: A Critical Feature of Bacterial Cell Replication
Abstract
Spatial organization is a hallmark of all living systems. Even bacteria, the smallest forms of cellular life, display defined shapes and complex internal organization, showcasing a highly structured genome, cytoskeletal filaments, localized scaffolding structures, dynamic spatial patterns, active transport, and occasionally, intracellular organelles. Spatial order is required for faithful and efficient cellular replication and offers a powerful means for the development of unique biological properties. Here, we discuss organizational features of bacterial cells and highlight how bacteria have evolved diverse spatial mechanisms to overcome challenges cells face as self-replicating entities.
Copyright © 2018 Elsevier Inc. All rights reserved.
Figures

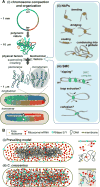



References
-
- Aaron M, Charbon G, Lam H, Schwarz H, Vollmer W, Jacobs-Wagner C. The tubulin homologue FtsZ contributes to cell elongation by guiding cell wall precursor synthesis in Caulobacter crescentus. Mol Microbiol. 2007;64:938–52. - PubMed
-
- Abdul-Rahman F, Petit E, Blanchard JL. Distribution of polyhedral bacterial microcompartments suggests frequent horizontal transfer and operon reassembly. J Phylogen Evolution Biol. 2013;1:118.
-
- Adachi S, Hori K, Hiraga S. Subcellular positioning of F plasmid mediated by dynamic localization of SopA and SopB. J Mol Biol. 2006;356:850–63. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

