Signature of circular RNAs in human induced pluripotent stem cells and derived cardiomyocytes
- PMID: 29523209
- PMCID: PMC5845222
- DOI: 10.1186/s13287-018-0793-5
Signature of circular RNAs in human induced pluripotent stem cells and derived cardiomyocytes
Abstract
Background: Circular RNAs (circRNAs) are regarded as a novel class of noncoding RNA regulators. Although a number of circRNAs have been identified by bioinformatics analysis of RNA-seq data, tissue and disease-specific circRNAs are still to be uncovered to promote their application in basic research and clinical practice. The purpose of this study was to explore the circRNA profiles in human induced pluripotent stem cells (hiPSCs) and hiPSC-derived cardiomyocytes (hiPSC-CMs), and to identify cardiac or disease-specific circRNAs.
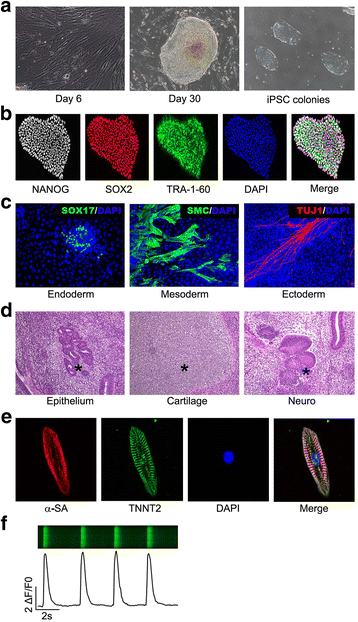
Methods: hiPSCs were generated from fibroblasts, and then further differentiated to hiPSC-CMs by modulating WNT signaling in RPMI+B27 medium. Following high-throughput RNA sequencing, circRNAs were extracted and quantified by a combined strategy known as CIRCexplorer. Integrative analysis was performed to illuminate the correlation between circRNAs and their parental linear isoforms. Cardiac and disease-specific expression of circRNAs was confirmed by quantitative reverse-transcription PCR.
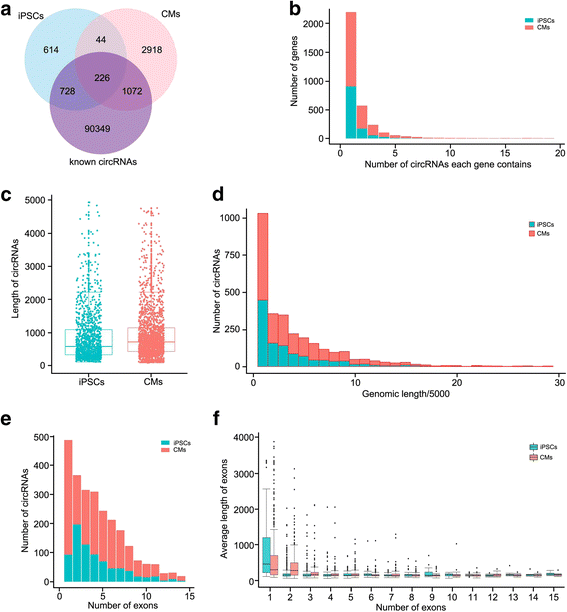
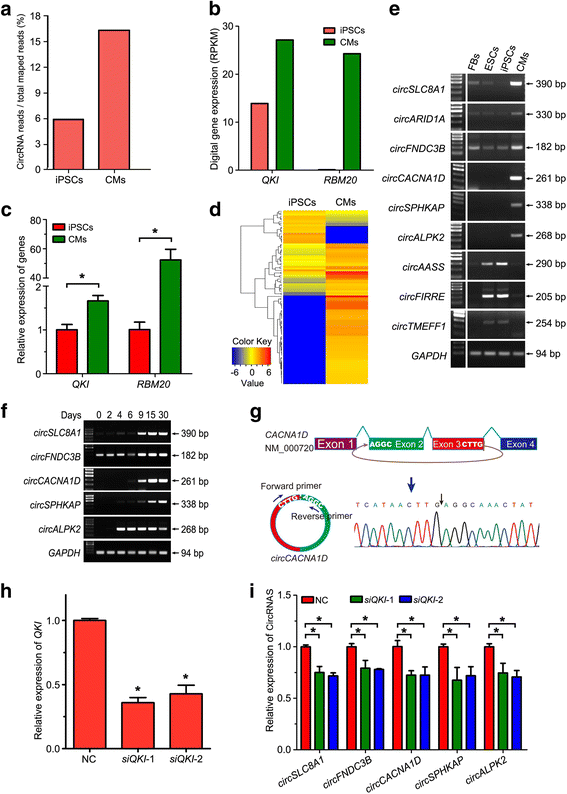
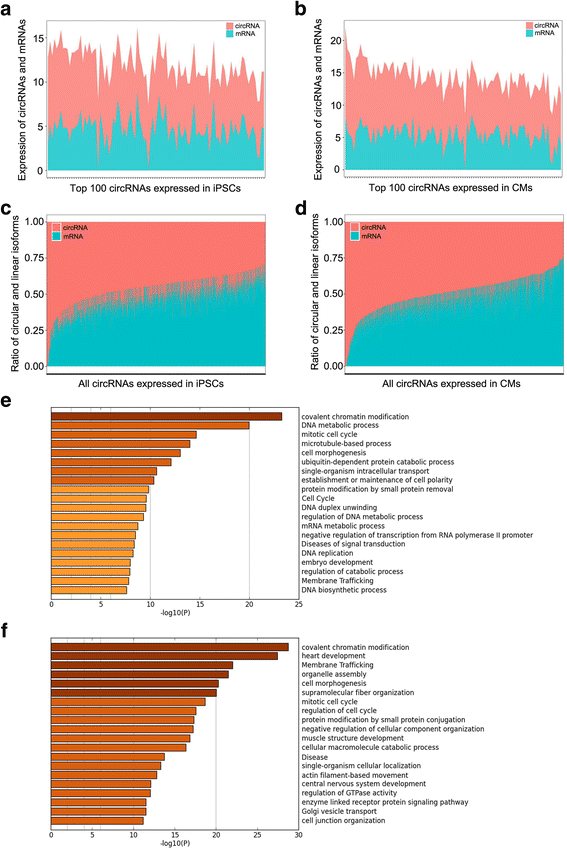
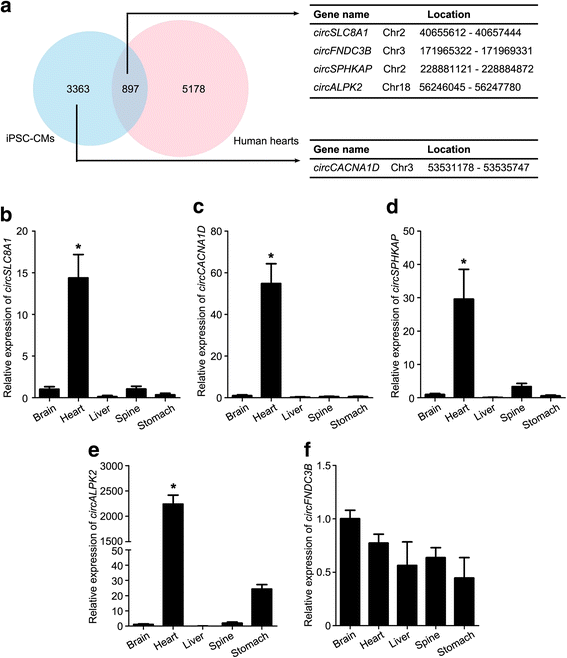
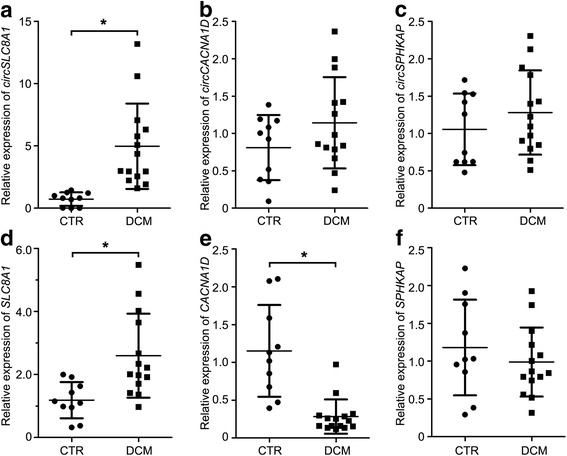
Results: In this study, a total of 5602 circRNAs were identified in hiPSCs and hiPSC-CMs. Our data indicated, for the first time, more enriched expression of circRNAs in differentiated cardiomyocytes than in undifferentiated hiPSCs. In addition to the host gene-dependent expression, our integrative analysis also identified a number of circRNAs showing host gene-independent expression in hiPSCs and hiPSC-CMs. CircRNAs including circSLC8A1, circCACNA1D, circSPHKAP and circALPK2 showed cardiac-selective expression during cardiac differentiation and human heart-specific enrichment in fetal tissues. Furthermore, circSLC8A1 abnormally increased in heart tissues from patients suffering from dilated cardiomyopathy.
Conclusions: CircRNAs are highly enriched in hiPSC-differentiated CMs, and cardiac-specific circRNAs such as circSLC8A1, circCACNA1D, circSPHKAP and circALPK2 may serve as biomarkers of CMs. Detection of the excessive expression of circSLC8A1 provides a potential approach for pathological status indication of heart disease.
Keywords: Cardiomyocyte; Circular RNA; Induced pluripotent stem cell.
Conflict of interest statement
Ethics approval and consent to participate
Experiments with donated human tissues including adult hearts and skin biopsies, as well as fetal brain, heart, liver, spine and stomach, were approved by the ethical committees of Soochow University, Suzhou, China and Huazhong University of Science and Technology, Wuhan, China, and performed after informed consent from the donors or their parents/legal guardians.
Consent for publication
All authors of this manuscript agreed to publication.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
Similar articles
-
Identification of circular RNAs with host gene-independent expression in human model systems for cardiac differentiation and disease.J Mol Cell Cardiol. 2017 Aug;109:48-56. doi: 10.1016/j.yjmcc.2017.06.015. Epub 2017 Jul 1. J Mol Cell Cardiol. 2017. PMID: 28676412
-
Deep Characterization of Circular RNAs from Human Cardiovascular Cell Models and Cardiac Tissue.Cells. 2020 Jul 4;9(7):1616. doi: 10.3390/cells9071616. Cells. 2020. PMID: 32635460 Free PMC article.
-
Human-induced pluripotent stem cell-derived cardiomyocytes from cardiac progenitor cells: effects of selective ion channel blockade.Europace. 2016 Dec;18(suppl 4):iv67-iv76. doi: 10.1093/europace/euw352. Europace. 2016. PMID: 28011833
-
Functions and mechanisms of circular RNAs in regulating stem cell differentiation.RNA Biol. 2021 Dec;18(12):2136-2149. doi: 10.1080/15476286.2021.1913551. Epub 2021 Apr 26. RNA Biol. 2021. PMID: 33896374 Free PMC article. Review.
-
Production of functional cardiomyocytes and cardiac tissue from human induced pluripotent stem cells for regenerative therapy.J Mol Cell Cardiol. 2022 Mar;164:83-91. doi: 10.1016/j.yjmcc.2021.11.008. Epub 2021 Nov 22. J Mol Cell Cardiol. 2022. PMID: 34822838 Review.
Cited by
-
Development and disease-specific regulation of RNA splicing in cardiovascular system.Front Cell Dev Biol. 2024 Jul 9;12:1423553. doi: 10.3389/fcell.2024.1423553. eCollection 2024. Front Cell Dev Biol. 2024. PMID: 39045460 Free PMC article. Review.
-
The emerging roles of circular RNAs in regulating the fate of stem cells.Mol Cell Biochem. 2021 Jan;476(1):231-246. doi: 10.1007/s11010-020-03900-w. Epub 2020 Sep 11. Mol Cell Biochem. 2021. PMID: 32918186 Review.
-
Establishment of an in vitro safety assessment model for lipid-lowering drugs using same-origin human pluripotent stem cell-derived cardiomyocytes and endothelial cells.Acta Pharmacol Sin. 2022 Jan;43(1):240-250. doi: 10.1038/s41401-021-00621-8. Epub 2021 Mar 8. Acta Pharmacol Sin. 2022. PMID: 33686244 Free PMC article.
-
Circular RNA Expression for Dilated Cardiomyopathy in Hearts and Pluripotent Stem Cell-Derived Cardiomyocytes.Front Cell Dev Biol. 2021 Dec 17;9:760515. doi: 10.3389/fcell.2021.760515. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34977015 Free PMC article. Review.
-
Generation of human vascularized and chambered cardiac organoids for cardiac disease modelling and drug evaluation.Cell Prolif. 2024 Aug;57(8):e13631. doi: 10.1111/cpr.13631. Epub 2024 Mar 7. Cell Prolif. 2024. PMID: 38453465 Free PMC article.
References
-
- Ye L, Chang YH, Xiong Q, Zhang P, Zhang L, Somasundaram P, Lepley M, Swingen C, Su L, Wendel JS, et al. Cardiac repair in a porcine model of acute myocardial infarction with human induced pluripotent stem cell-derived cardiovascular cells. Cell Stem Cell. 2014;15:750–761. doi: 10.1016/j.stem.2014.11.009. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

