Folding-Degradation Relationship of a Membrane Protein Mediated by the Universally Conserved ATP-Dependent Protease FtsH
- PMID: 29528632
- PMCID: PMC5892824
- DOI: 10.1021/jacs.8b00832
Folding-Degradation Relationship of a Membrane Protein Mediated by the Universally Conserved ATP-Dependent Protease FtsH
Abstract
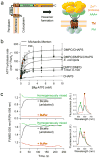
ATP-dependent protein degradation mediated by AAA+ proteases is one of the major cellular pathways for protein quality control and regulation of functional networks. While a majority of studies of protein degradation have focused on water-soluble proteins, it is not well understood how membrane proteins with abnormal conformation are selectively degraded. The knowledge gap stems from the lack of an in vitro system in which detailed molecular mechanisms can be studied as well as difficulties in studying membrane protein folding in lipid bilayers. To quantitatively define the folding-degradation relationship of membrane proteins, we reconstituted the degradation using the conserved membrane-integrated AAA+ protease FtsH as a model degradation machine and the stable helical-bundle membrane protein GlpG as a model substrate in the lipid bilayer environment. We demonstrate that FtsH possesses a substantial ability to actively unfold GlpG, and the degradation significantly depends on the stability and hydrophobicity near the degradation marker. We find that FtsH hydrolyzes 380-550 ATP molecules to degrade one copy of GlpG. Remarkably, FtsH overcomes the dual-energetic burden of substrate unfolding and membrane dislocation with the ATP cost comparable to that for water-soluble substrates by robust ClpAP/XP proteases. The physical principles elucidated in this study provide general insights into membrane protein degradation mediated by ATP-dependent proteolytic systems.
Figures


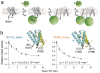

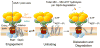
Similar articles
-
Proteolysis mediated by the membrane-integrated ATP-dependent protease FtsH has a unique nonlinear dependence on ATP hydrolysis rates.Protein Sci. 2019 Jul;28(7):1262-1275. doi: 10.1002/pro.3629. Epub 2019 May 8. Protein Sci. 2019. PMID: 31008538 Free PMC article.
-
Structure and function of the bacterial AAA protease FtsH.Biochim Biophys Acta. 2012 Jan;1823(1):40-8. doi: 10.1016/j.bbamcr.2011.08.015. Epub 2011 Sep 8. Biochim Biophys Acta. 2012. PMID: 21925212 Review.
-
Reconstitution and functional characterization of the FtsH protease in lipid nanodiscs.Biochim Biophys Acta Biomembr. 2021 Feb 1;1863(2):183526. doi: 10.1016/j.bbamem.2020.183526. Epub 2020 Dec 2. Biochim Biophys Acta Biomembr. 2021. PMID: 33278347
-
When, how and why? Regulated proteolysis by the essential FtsH protease in Escherichia coli.Biol Chem. 2017 May 1;398(5-6):625-635. doi: 10.1515/hsz-2016-0302. Biol Chem. 2017. PMID: 28085670 Review.
-
Structural insights into the membrane-bound proteolytic machinery of bacterial protein quality control.Biochem Soc Trans. 2024 Oct 30;52(5):2077-2086. doi: 10.1042/BST20231250. Biochem Soc Trans. 2024. PMID: 39417347 Review.
Cited by
-
Untangling the complexity of membrane protein folding.Curr Opin Struct Biol. 2022 Feb;72:237-247. doi: 10.1016/j.sbi.2021.11.013. Epub 2022 Jan 5. Curr Opin Struct Biol. 2022. PMID: 34995926 Free PMC article. Review.
-
GeTFEP: A general transfer free energy profile of transmembrane proteins.Protein Sci. 2020 Feb;29(2):469-479. doi: 10.1002/pro.3763. Epub 2019 Nov 11. Protein Sci. 2020. PMID: 31658402 Free PMC article.
-
On the Interpretation of Force-Induced Unfolding Studies of Membrane Proteins Using Fast Simulations.Biophys J. 2019 Oct 15;117(8):1429-1441. doi: 10.1016/j.bpj.2019.09.011. Epub 2019 Sep 17. Biophys J. 2019. PMID: 31587831 Free PMC article.
-
The AAA + ATPase valosin-containing protein (VCP)/p97/Cdc48 interaction network in Leishmania.Sci Rep. 2020 Aug 4;10(1):13135. doi: 10.1038/s41598-020-70010-4. Sci Rep. 2020. PMID: 32753747 Free PMC article.
-
Lipid bilayer induces contraction of the denatured state ensemble of a helical-bundle membrane protein.Proc Natl Acad Sci U S A. 2022 Jan 4;119(1):e2109169119. doi: 10.1073/pnas.2109169119. Proc Natl Acad Sci U S A. 2022. PMID: 34969836 Free PMC article.
References
-
- Gottesman S. Proteases and their targets in Escherichia coli. Annu Rev Genet. 1996;30:465–506. - PubMed
-
- Sauer RT, Baker TA. AAA+ proteases: ATP-fueled machines of protein destruction. Annu Rev Biochem. 2011;80:587–612. - PubMed
-
- Langer T. AAA proteases: cellular machines for degrading membrane proteins. Trends Biochem Sci. 2000;25(5):247–251. - PubMed
-
- Ito K, Akiyama Y. Cellular functions, mechanism of action, and regulation of FtsH protease. Annu Rev Microbiol. 2005;59:211–231. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

