Metabolic Heterogeneity Evidenced by MRS among Patient-Derived Glioblastoma Multiforme Stem-Like Cells Accounts for Cell Clustering and Different Responses to Drugs
- PMID: 29531533
- PMCID: PMC5835274
- DOI: 10.1155/2018/3292704
Metabolic Heterogeneity Evidenced by MRS among Patient-Derived Glioblastoma Multiforme Stem-Like Cells Accounts for Cell Clustering and Different Responses to Drugs
Abstract
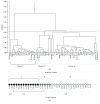
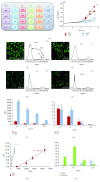
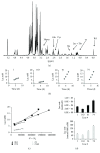
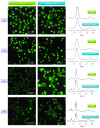
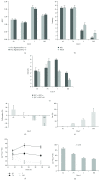
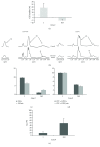
Clustering of patient-derived glioma stem-like cells (GSCs) through unsupervised analysis of metabolites detected by magnetic resonance spectroscopy (MRS) evidenced three subgroups, namely clusters 1a and 1b, with high intergroup similarity and neural fingerprints, and cluster 2, with a metabolism typical of commercial tumor lines. In addition, subclones generated by the same GSC line showed different metabolic phenotypes. Aerobic glycolysis prevailed in cluster 2 cells as demonstrated by higher lactate production compared to cluster 1 cells. Oligomycin, a mitochondrial ATPase inhibitor, induced high lactate extrusion only in cluster 1 cells, where it produced neutral lipid accumulation detected as mobile lipid signals by MRS and lipid droplets by confocal microscopy. These results indicate a relevant role of mitochondrial fatty acid oxidation for energy production in GSCs. On the other hand, further metabolic differences, likely accounting for different therapy responsiveness observed after etomoxir treatment, suggest that caution must be used in considering patient treatment with mitochondria FAO blockers. Metabolomics and metabolic profiling may contribute to discover new diagnostic or prognostic biomarkers to be used for personalized therapies.
Figures

References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

