Ube2V2 Is a Rosetta Stone Bridging Redox and Ubiquitin Codes, Coordinating DNA Damage Responses
- PMID: 29532025
- PMCID: PMC5833000
- DOI: 10.1021/acscentsci.7b00556
Ube2V2 Is a Rosetta Stone Bridging Redox and Ubiquitin Codes, Coordinating DNA Damage Responses
Abstract
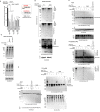
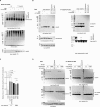
Posttranslational modifications (PTMs) are the lingua franca of cellular communication. Most PTMs are enzyme-orchestrated. However, the reemergence of electrophilic drugs has ushered mining of unconventional/non-enzyme-catalyzed electrophile-signaling pathways. Despite the latest impetus toward harnessing kinetically and functionally privileged cysteines for electrophilic drug design, identifying these sensors remains challenging. Herein, we designed "G-REX"-a technique that allows controlled release of reactive electrophiles in vivo. Mitigating toxicity/off-target effects associated with uncontrolled bolus exposure, G-REX tagged first-responding innate cysteines that bind electrophiles under true kcat/Km conditions. G-REX identified two allosteric ubiquitin-conjugating proteins-Ube2V1/Ube2V2-sharing a novel privileged-sensor-cysteine. This non-enzyme-catalyzed-PTM triggered responses specific to each protein. Thus, G-REX is an unbiased method to identify novel functional cysteines. Contrasting conventional active-site/off-active-site cysteine-modifications that regulate target activity, modification of Ube2V2 allosterically hyperactivated its enzymatically active binding-partner Ube2N, promoting K63-linked client ubiquitination and stimulating H2AX-dependent DNA damage response. This work establishes Ube2V2 as a Rosetta-stone bridging redox and ubiquitin codes to guard genome integrity.
Conflict of interest statement
The authors declare the following competing financial interest(s): Provisional patent for the G-REX method in filing stages (by Cornell University).
Figures



Similar articles
-
REX technologies for profiling and decoding the electrophile signaling axes mediated by Rosetta Stone proteins.Methods Enzymol. 2020;633:203-230. doi: 10.1016/bs.mie.2019.02.039. Epub 2019 Mar 14. Methods Enzymol. 2020. PMID: 32046846 Free PMC article.
-
An Oculus to Profile and Probe Target Engagement In Vivo: How T-REX Was Born and Its Evolution into G-REX.Acc Chem Res. 2021 Feb 2;54(3):618-631. doi: 10.1021/acs.accounts.0c00537. Epub 2020 Nov 23. Acc Chem Res. 2021. PMID: 33228351
-
Interrogating Precision Electrophile Signaling.Trends Biochem Sci. 2019 Apr;44(4):380-381. doi: 10.1016/j.tibs.2019.01.006. Epub 2019 Feb 11. Trends Biochem Sci. 2019. PMID: 30765181 Free PMC article.
-
Privileged Electrophile Sensors: A Resource for Covalent Drug Development.Cell Chem Biol. 2017 Jul 20;24(7):787-800. doi: 10.1016/j.chembiol.2017.05.023. Epub 2017 Jun 22. Cell Chem Biol. 2017. PMID: 28648380 Free PMC article. Review.
-
Chemical-proteomic strategies to investigate cysteine posttranslational modifications.Biochim Biophys Acta. 2014 Dec;1844(12):2315-30. doi: 10.1016/j.bbapap.2014.09.024. Epub 2014 Oct 5. Biochim Biophys Acta. 2014. PMID: 25291386 Review.
Cited by
-
A primer on harnessing non-enzymatic post-translational modifications for drug design.RSC Med Chem. 2021 Oct 26;12(11):1797-1807. doi: 10.1039/d1md00157d. eCollection 2021 Nov 17. RSC Med Chem. 2021. PMID: 34825181 Free PMC article.
-
Function-guided proximity mapping unveils electrophilic-metabolite sensing by proteins not present in their canonical locales.Proc Natl Acad Sci U S A. 2022 Feb 1;119(5):e2120687119. doi: 10.1073/pnas.2120687119. Proc Natl Acad Sci U S A. 2022. PMID: 35082156 Free PMC article.
-
Z-REX uncovers a bifurcation in function of Keap1 paralogs.Elife. 2022 Oct 27;11:e83373. doi: 10.7554/eLife.83373. Elife. 2022. PMID: 36300632 Free PMC article.
-
Genie in a bottle: controlled release helps tame natural polypharmacology?Curr Opin Chem Biol. 2019 Aug;51:48-56. doi: 10.1016/j.cbpa.2019.02.014. Epub 2019 Mar 23. Curr Opin Chem Biol. 2019. PMID: 30913473 Free PMC article. Review.
-
Breaking the Fourth Wall: Modulating Quaternary Associations for Protein Regulation and Drug Discovery.Chembiochem. 2019 May 2;20(9):1091-1104. doi: 10.1002/cbic.201800716. Epub 2019 Apr 1. Chembiochem. 2019. PMID: 30589188 Free PMC article. Review.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

