Genomic and Genotypic Characterization of Cylindrospermopsis raciborskii: Toward an Intraspecific Phylogenetic Evaluation by Comparative Genomics
- PMID: 29535689
- PMCID: PMC5834425
- DOI: 10.3389/fmicb.2018.00306
Genomic and Genotypic Characterization of Cylindrospermopsis raciborskii: Toward an Intraspecific Phylogenetic Evaluation by Comparative Genomics
Erratum in
-
Corrigendum: Genomic and Genotypic Characterization of Cylindrospermopsis raciborskii: Toward an Intraspecific Phylogenetic Evaluation by Comparative Genomics.Front Microbiol. 2018 May 15;9:979. doi: 10.3389/fmicb.2018.00979. eCollection 2018. Front Microbiol. 2018. PMID: 29795803 Free PMC article.
Abstract
Cylindrospermopsis raciborskii is a freshwater cyanobacterial species with increasing bloom reports worldwide that are likely due to factors related to climate change. In addition to the deleterious effects of blooms on aquatic ecosystems, the majority of ecotypes can synthesize toxic secondary metabolites causing public health issues. To overcome the harmful effects of C. raciborskii blooms, it is important to advance knowledge of diversity, genetic variation, and evolutionary processes within populations. An efficient approach to exploring this diversity and understanding the evolution of C. raciborskii is to use comparative genomics. Here, we report two new draft genomes of C. raciborskii (strains CENA302 and CENA303) from Brazilian isolates of different origins and explore their molecular diversity, phylogeny, and evolutionary diversification by comparing their genomes with sequences from other strains available in public databases. The results obtained by comparing seven C. raciborskii and the Raphidiopsis brookii D9 genomes revealed a set of conserved core genes and a variable set of accessory genes, such as those involved in the biosynthesis of natural products, heterocyte glycolipid formation, and nitrogen fixation. Gene cluster arrangements related to the biosynthesis of the antifungal cyclic glycosylated lipopeptide hassallidin were identified in four C. raciborskii genomes, including the non-nitrogen fixing strain CENA303. Shifts in gene clusters involved in toxin production according to geographic origins were observed, as well as a lack of nitrogen fixation (nif) and heterocyte glycolipid (hgl) gene clusters in some strains. Single gene phylogeny (16S rRNA sequences) was congruent with phylogeny based on 31 concatenated housekeeping protein sequences, and both analyses have shown, with high support values, that the species C. raciborskii is monophyletic. This comparative genomics study allowed a species-wide view of the biological diversity of C. raciborskii and in some cases linked genome differences to phenotype.
Keywords: bioinformatics; cyanobacteria; cyanotoxins; genome assembly; natural products; nitrogen fixation; pan-genome.
Figures
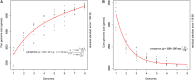




References
-
- Aguillera A., Berrendero Gómez E., Kaštovský J., Echenique R. O., Salerno G. (2018). The polyphasic analysis of two native Raphidiopsis isolates supports the unification of the genera Raphidiopsis and Cylindrospermopsis (Nostocales, Cyanobacteria). 57 130–146. 10.2216/17-2.1 - DOI
-
- Alster A., Kaplan-Levy R. N., Sukenik A., Zohary T. (2010). Morphology and phylogeny of a non-toxic invasive Cylindrospermopsis raciborskii from a Mediterranean Lake. 639 115–128. 10.1007/s10750-009-0044-y - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources

