Target identification reveals protein arginine methyltransferase 1 is a potential target of phenyl vinyl sulfone and its derivatives
- PMID: 29540535
- PMCID: PMC5968187
- DOI: 10.1042/BSR20171717
Target identification reveals protein arginine methyltransferase 1 is a potential target of phenyl vinyl sulfone and its derivatives
Abstract
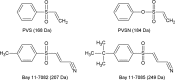
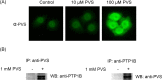
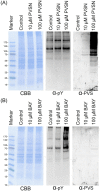
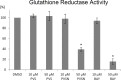
Phenyl vinyl sulfone (PVS) and phenyl vinyl sulfonate (PVSN) inactivate protein tyrosine phosphatases (PTPs) by mimicking the phosphotyrosine structure and providing a Michael addition acceptor for the active-site cysteine residue of PTPs, thus forming covalent adducts between PVS (or PVSN) and PTPs. We developed a specific antiserum against PVS. This antiserum can be used in general antibody-based assays such as immunoblotting, immunofluorescence staining, and immunoprecipitation. Target identification through immunoprecipitation and mass spectrometry analysis reveals potential targets of PVS, mostly proteins with reactive cysteine residues or low-pKa cysteine residues that are prone to reversible redox modifications. Target identification of PVSN has been conducted because the anti-PVS antiserum can also recognize PVSN. Among the targets, protein arginine methyltransferase 1 (PRMT1), inosine-5'-monophosphate dehydrogenase 1, vimentin, and glutathione reductase (GR) were further confirmed by immunoprecipitation followed by immunoblotting. In addition, PVSN and Bay11-7082 inhibited GR activity, and PVS, PVSN, and Bay 11-7082 inhibited PRMT1 activity in in vitro assays. In addition, treatment of PVSN, Bay11-7082, or Bay 11-7085 in cultured HeLa cells can cause the quick decline in the levels of protein asymmetric dimethylarginine. These results indicate that the similar moiety among PVS, PVSN, Bay 11-7082, and Bay 11-7085 can be the key structure of lead compounds of PRMT1. Therefore, we expect to use this approach in the identification of potential targets of other covalent drugs.
Keywords: Bay11-7082; covalent drug; phenyl vinyl sulfonate; phenyl vinyl sulfone; protein arginine methyltransferase 1; target identification.
© 2018 The Author(s).
Conflict of interest statement
The authors declare that there are no competing interests associated with the manuscript.
Figures






Similar articles
-
Aryl vinyl sulfonates and sulfones as active site-directed and mechanism-based probes for protein tyrosine phosphatases.J Am Chem Soc. 2008 Jul 2;130(26):8251-60. doi: 10.1021/ja711125p. Epub 2008 Jun 4. J Am Chem Soc. 2008. PMID: 18528979 Free PMC article.
-
Redox Control of Protein Arginine Methyltransferase 1 (PRMT1) Activity.J Biol Chem. 2015 Jun 12;290(24):14915-26. doi: 10.1074/jbc.M115.651380. Epub 2015 Apr 24. J Biol Chem. 2015. PMID: 25911106 Free PMC article.
-
A protein arginine N-methyltransferase 1 (PRMT1) and 2 heteromeric interaction increases PRMT1 enzymatic activity.Biochemistry. 2011 Sep 27;50(38):8226-40. doi: 10.1021/bi200644c. Epub 2011 Aug 29. Biochemistry. 2011. PMID: 21851090
-
The physiological and pathophysiological role of PRMT1-mediated protein arginine methylation.Pharmacol Res. 2009 Dec;60(6):466-74. doi: 10.1016/j.phrs.2009.07.006. Epub 2009 Jul 28. Pharmacol Res. 2009. PMID: 19643181 Review.
-
The vinyl sulfone motif as a structural unit for novel drug design and discovery.Expert Opin Drug Discov. 2024 Feb;19(2):239-251. doi: 10.1080/17460441.2023.2284201. Epub 2024 Feb 1. Expert Opin Drug Discov. 2024. PMID: 37978948 Review.
Cited by
-
A Genome-wide Haploid Genetic Screen Identifies Regulators of Glutathione Abundance and Ferroptosis Sensitivity.Cell Rep. 2019 Feb 5;26(6):1544-1556.e8. doi: 10.1016/j.celrep.2019.01.043. Cell Rep. 2019. PMID: 30726737 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

