Xanthomonas citri jumbo phage XacN1 exhibits a wide host range and high complement of tRNA genes
- PMID: 29540765
- PMCID: PMC5852040
- DOI: 10.1038/s41598-018-22239-3
Xanthomonas citri jumbo phage XacN1 exhibits a wide host range and high complement of tRNA genes
Abstract
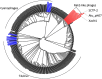
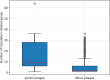
Xanthomonas virus (phage) XacN1 is a novel jumbo myovirus infecting Xanthomonas citri, the causative agent of Asian citrus canker. Its linear 384,670 bp double-stranded DNA genome encodes 592 proteins and presents the longest (66 kbp) direct terminal repeats (DTRs) among sequenced viral genomes. The DTRs harbor 56 tRNA genes, which correspond to all 20 amino acids and represent the largest number of tRNA genes reported in a viral genome. Codon usage analysis revealed a propensity for the phage encoded tRNAs to target codons that are highly used by the phage but less frequently by its host. The existence of these tRNA genes and seven additional translation-related genes as well as a chaperonin gene found in the XacN1 genome suggests a relative independence of phage replication on host molecular machinery, leading to a prediction of a wide host range for this jumbo phage. We confirmed the prediction by showing a wider host range of XacN1 than other X. citri phages in an infection test against a panel of host strains. Phylogenetic analyses revealed a clade of phages composed of XacN1 and ten other jumbo phages, indicating an evolutionary stable large genome size for this group of phages.
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
Xanthomonas bacteriophages: a review of their biology and biocontrol applications in agriculture.BMC Microbiol. 2021 Oct 25;21(1):291. doi: 10.1186/s12866-021-02351-7. BMC Microbiol. 2021. PMID: 34696726 Free PMC article. Review.
-
Intracellular Organization by Jumbo Bacteriophages.J Bacteriol. 2020 Dec 18;203(2):e00362-20. doi: 10.1128/JB.00362-20. Print 2020 Dec 18. J Bacteriol. 2020. PMID: 32868402 Free PMC article. Review.
-
Isolation and characterization of vB_XciM_LucasX, a new jumbo phage that infects Xanthomonas citri and Xanthomonas fuscans.PLoS One. 2022 Apr 14;17(4):e0266891. doi: 10.1371/journal.pone.0266891. eCollection 2022. PLoS One. 2022. PMID: 35421196 Free PMC article.
-
Bioinformatic analysis of the Acinetobacter baumannii phage AB1 genome.Gene. 2012 Oct 10;507(2):125-34. doi: 10.1016/j.gene.2012.07.029. Epub 2012 Jul 31. Gene. 2012. PMID: 22868206
-
Characterization of Novel Erwinia amylovora Jumbo Bacteriophages from Eneladusvirus Genus.Viruses. 2020 Nov 30;12(12):1373. doi: 10.3390/v12121373. Viruses. 2020. PMID: 33266226 Free PMC article.
Cited by
-
Xanthomonas bacteriophages: a review of their biology and biocontrol applications in agriculture.BMC Microbiol. 2021 Oct 25;21(1):291. doi: 10.1186/s12866-021-02351-7. BMC Microbiol. 2021. PMID: 34696726 Free PMC article. Review.
-
A persistent giant algal virus, with a unique morphology, encodes an unprecedented number of genes involved in energy metabolism.J Virol. 2021 Mar 25;95(8):e02446-20. doi: 10.1128/JVI.02446-20. Epub 2021 Feb 3. J Virol. 2021. PMID: 33536167 Free PMC article.
-
Codon usage revisited: Lack of correlation between codon usage and the number of tRNA genes in enterobacteria.Biochem Biophys Res Commun. 2018 Aug 25;502(4):450-455. doi: 10.1016/j.bbrc.2018.05.168. Epub 2018 Jun 5. Biochem Biophys Res Commun. 2018. PMID: 29859934 Free PMC article.
-
Isolation and Characterization of Two Novel Genera of Jumbo Bacteriophages Infecting Xanthomonas vesicatoria Isolated from Agricultural Regions in Mexico.Antibiotics (Basel). 2024 Jul 15;13(7):651. doi: 10.3390/antibiotics13070651. Antibiotics (Basel). 2024. PMID: 39061333 Free PMC article.
-
Intracellular Organization by Jumbo Bacteriophages.J Bacteriol. 2020 Dec 18;203(2):e00362-20. doi: 10.1128/JB.00362-20. Print 2020 Dec 18. J Bacteriol. 2020. PMID: 32868402 Free PMC article. Review.
References
-
- Hendrix RW. Jumbo bacteriophages. Curr. Top. Microbiol. Immunol. 2009;328:229–240. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

