How RNA transcripts coordinate DNA recombination and repair
- PMID: 29545568
- PMCID: PMC5854605
- DOI: 10.1038/s41467-018-03483-7
How RNA transcripts coordinate DNA recombination and repair
Abstract
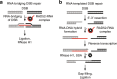
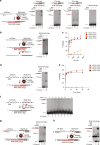
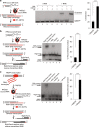
Genetic studies in yeast indicate that RNA transcripts facilitate homology-directed DNA repair in a manner that is dependent on RAD52. The molecular basis for so-called RNA-DNA repair, however, remains unknown. Using reconstitution assays, we demonstrate that RAD52 directly cooperates with RNA as a sequence-directed ribonucleoprotein complex to promote two related modes of RNA-DNA repair. In a RNA-bridging mechanism, RAD52 assembles recombinant RNA-DNA hybrids that coordinate synapsis and ligation of homologous DNA breaks. In an RNA-templated mechanism, RAD52-mediated RNA-DNA hybrids enable reverse transcription-dependent RNA-to-DNA sequence transfer at DNA breaks that licenses subsequent DNA recombination. Notably, we show that both mechanisms of RNA-DNA repair are promoted by transcription of a homologous DNA template in trans. In summary, these data elucidate how RNA transcripts cooperate with RAD52 to coordinate homology-directed DNA recombination and repair in the absence of a DNA donor, and demonstrate a direct role for transcription in RNA-DNA repair.
Conflict of interest statement
The authors declare no competing interests.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

