CDK and MAPK Synergistically Regulate Signaling Dynamics via a Shared Multi-site Phosphorylation Region on the Scaffold Protein Ste5
- PMID: 29547722
- PMCID: PMC5858200
- DOI: 10.1016/j.molcel.2018.02.018
CDK and MAPK Synergistically Regulate Signaling Dynamics via a Shared Multi-site Phosphorylation Region on the Scaffold Protein Ste5
Abstract
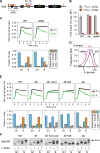
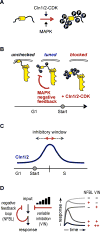
We report an unanticipated system of joint regulation by cyclin-dependent kinase (CDK) and mitogen-activated protein kinase (MAPK), involving collaborative multi-site phosphorylation of a single substrate. In budding yeast, the protein Ste5 controls signaling through a G1 arrest pathway. Upon cell-cycle entry, CDK inhibits Ste5 via multiple phosphorylation sites, disrupting its membrane association. Using quantitative time-lapse microscopy, we examined Ste5 membrane recruitment dynamics at different cell-cycle stages. Surprisingly, in S phase, where Ste5 recruitment should be blocked, we observed an initial recruitment followed by a steep drop-off. This delayed inhibition revealed a requirement for both CDK activity and negative feedback from the pathway MAPK Fus3. Mutagenesis, mass spectrometry, and electrophoretic analyses suggest that the CDK and MAPK modify shared sites, which are most extensively phosphorylated when both kinases are active and able to bind their docking sites on Ste5. Such collaborative phosphorylation can broaden regulatory inputs and diversify output dynamics of signaling pathways.
Keywords: Cdc28; Cks1; Cln2; G protein; Ste4; cyclin; mating; pheromone; signal transduction; start.
Copyright © 2018 Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
Counteractive control of polarized morphogenesis during mating by mitogen-activated protein kinase Fus3 and G1 cyclin-dependent kinase.Mol Biol Cell. 2008 Apr;19(4):1739-52. doi: 10.1091/mbc.e07-08-0757. Epub 2008 Feb 6. Mol Biol Cell. 2008. PMID: 18256288 Free PMC article.
-
A mechanism for cell-cycle regulation of MAP kinase signaling in a yeast differentiation pathway.Cell. 2007 Feb 9;128(3):519-31. doi: 10.1016/j.cell.2006.12.032. Cell. 2007. PMID: 17289571 Free PMC article.
-
Cyclin-specific docking motifs promote phosphorylation of yeast signaling proteins by G1/S Cdk complexes.Curr Biol. 2011 Oct 11;21(19):1615-23. doi: 10.1016/j.cub.2011.08.033. Epub 2011 Sep 22. Curr Biol. 2011. PMID: 21945277 Free PMC article.
-
CDK signaling via nonconventional CDK phosphorylation sites.Mol Biol Cell. 2023 Nov 1;34(12):pe5. doi: 10.1091/mbc.E22-06-0196. Mol Biol Cell. 2023. PMID: 37906435 Free PMC article. Review.
-
Quantitative model of eukaryotic Cdk control through the Forkhead CONTROLLER.NPJ Syst Biol Appl. 2021 Jun 11;7(1):28. doi: 10.1038/s41540-021-00187-5. NPJ Syst Biol Appl. 2021. PMID: 34117265 Free PMC article. Review.
Cited by
-
Scaffold Proteins in Fibrotic Diseases of Visceral Organs.Biomolecules. 2025 Mar 16;15(3):420. doi: 10.3390/biom15030420. Biomolecules. 2025. PMID: 40149956 Free PMC article. Review.
-
Novel interconnections of HOG signaling revealed by combined use of two proteomic software packages.Cell Commun Signal. 2019 Jun 17;17(1):66. doi: 10.1186/s12964-019-0381-z. Cell Commun Signal. 2019. PMID: 31208443 Free PMC article.
-
Mechanical stress impairs pheromone signaling via Pkc1-mediated regulation of the MAPK scaffold Ste5.J Cell Biol. 2019 Sep 2;218(9):3117-3133. doi: 10.1083/jcb.201808161. Epub 2019 Jul 17. J Cell Biol. 2019. PMID: 31315942 Free PMC article.
-
Hyperosmolality in CHO cell culture: effects on the proteome.Appl Microbiol Biotechnol. 2022 Apr;106(7):2569-2586. doi: 10.1007/s00253-022-11861-x. Epub 2022 Mar 21. Appl Microbiol Biotechnol. 2022. PMID: 35312825 Free PMC article.
-
PROBABILISTIC LEARNING OF TREATMENT TREES IN CANCER.Ann Appl Stat. 2023 Sep;17(3):1884-1908. doi: 10.1214/22-aoas1696. Epub 2023 Sep 7. Ann Appl Stat. 2023. PMID: 37711665 Free PMC article.
References
-
- Alon U. An Introduction to Systems Biology: Design Principles of Biological Circuits. Chapman & Hall/CRC Mathematical & Computational Biology; Chapman & Hall; 2006.
-
- Bhattacharyya RP, Remenyi A, Good MC, Bashor CJ, Falick AM, Lim WA. The Ste5 scaffold allosterically modulates signaling output of the yeast mating pathway. Science. 2006;311:822–826. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

