Modulation of nucleosomal DNA accessibility via charge-altering post-translational modifications in histone core
- PMID: 29548294
- PMCID: PMC5856334
- DOI: 10.1186/s13072-018-0181-5
Modulation of nucleosomal DNA accessibility via charge-altering post-translational modifications in histone core
Abstract
Background: Controlled modulation of nucleosomal DNA accessibility via post-translational modifications (PTM) is a critical component to many cellular functions. Charge-altering PTMs in the globular histone core-including acetylation, phosphorylation, crotonylation, propionylation, butyrylation, formylation, and citrullination-can alter the strong electrostatic interactions between the oppositely charged nucleosomal DNA and the histone proteins and thus modulate accessibility of the nucleosomal DNA, affecting processes that depend on access to the genetic information, such as transcription. However, direct experimental investigation of the effects of these PTMs is very difficult. Theoretical models can rationalize existing observations, suggest working hypotheses for future experiments, and provide a unifying framework for connecting PTMs with the observed effects.
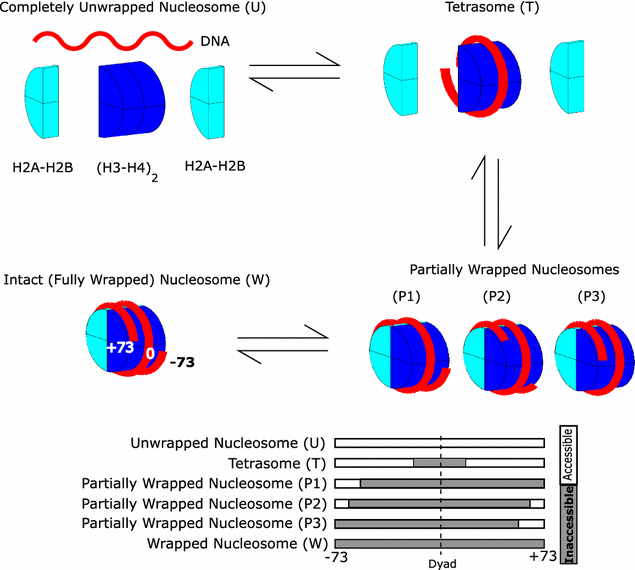
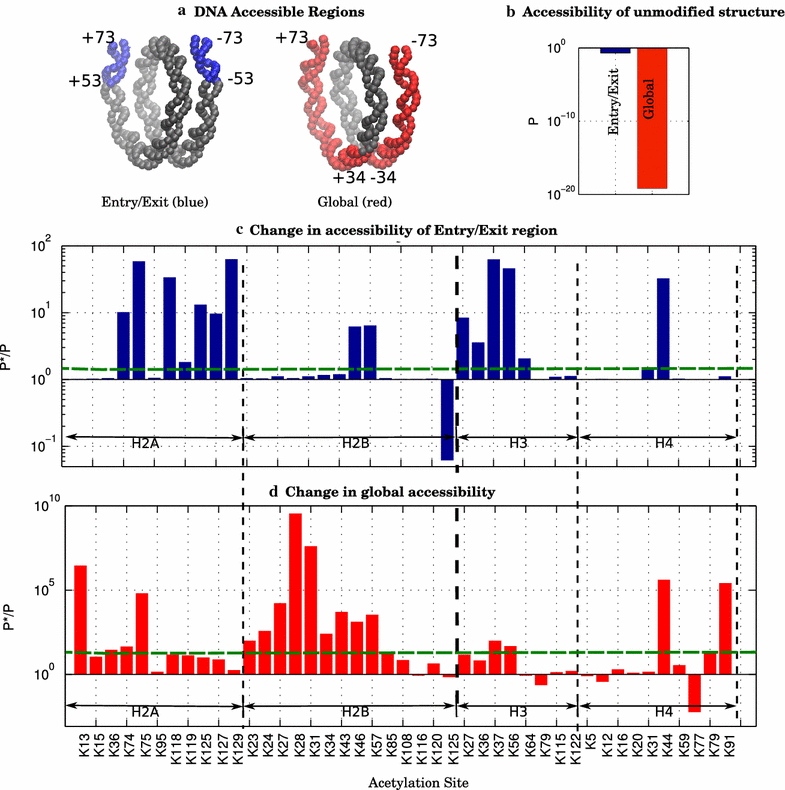
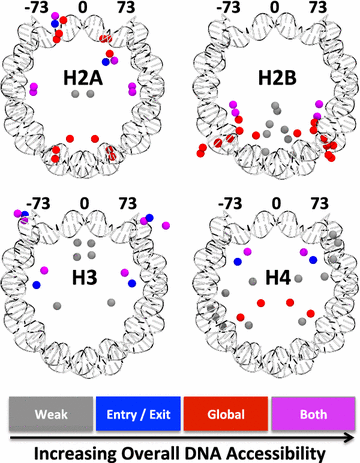
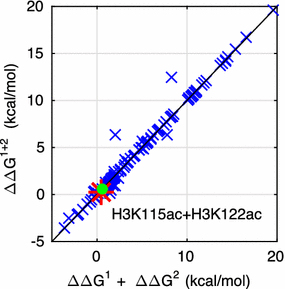
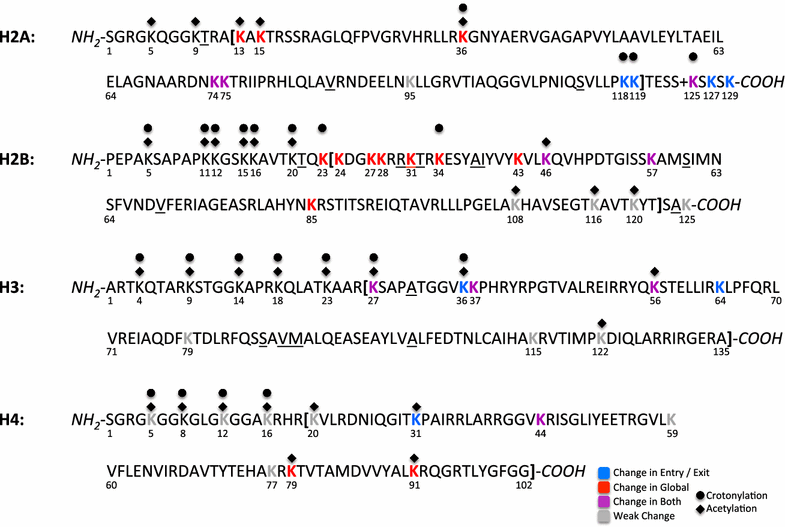
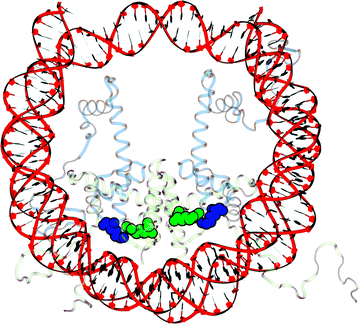
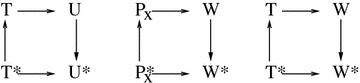
Results: A physics-based framework is proposed that predicts the effect of charge-altering PTMs in the histone core, quantitatively for several types of lysine charge-neutralizing PTMs including acetylation, and qualitatively for all phosphorylations, on the nucleosome stability and subsequent changes in DNA accessibility, making a connection to resulting biological phenotypes. The framework takes into account multiple partially assembled states of the nucleosome at the atomic resolution. The framework is validated against experimentally known nucleosome stability changes due to the acetylation of specific lysines, and their effect on transcription. The predicted effect of charge-altering PTMs on DNA accessibility can vary dramatically, from virtually none to a strong, region-dependent increase in accessibility of the nucleosomal DNA; in some cases, e.g., H4K44, H2AK75, and H2BK57, the effect is significantly stronger than that of the extensively studied acetylation sites such H3K56, H3K115 or H3K122. Proximity to the DNA is suggestive of the strength of the PTM effect, but there are many exceptions. For the vast majority of charge-altering PTMs, the predicted increase in the DNA accessibility should be large enough to result in a measurable modulation of transcription. However, a few possible PTMs, such as acetylation of H4K77, counterintuitively decrease the DNA accessibility, suggestive of the repressed chromatin. A structural explanation for the phenomenon is provided. For the majority of charge-altering PTMs, the effect on DNA accessibility is simply additive (noncooperative), but there are exceptions, e.g., simultaneous acetylation of H4K79 and H3K122, where the combined effect is amplified. The amplification is a direct consequence of the nucleosome-DNA complex having more than two structural states. The effect of individual PTMs is classified based on changes in the accessibility of various regions throughout the nucleosomal DNA. The PTM's resulting imprint on the DNA accessibility, "PTMprint," is used to predict effects of many yet unexplored PTMs. For example, acetylation of H4K44 yields a PTMprint similar to the PTMprint of H3K56, and thus acetylation of H4K44 is predicted to lead to a wide range of strong biological effects.
Conclusion: Charge-altering post-translational modifications in the relatively unexplored globular histone core may provide a precision mechanism for controlling accessibility to the nucleosomal DNA.
Figures
Similar articles
-
Histone fold modifications control nucleosome unwrapping and disassembly.Proc Natl Acad Sci U S A. 2011 Aug 2;108(31):12711-6. doi: 10.1073/pnas.1106264108. Epub 2011 Jul 18. Proc Natl Acad Sci U S A. 2011. PMID: 21768347 Free PMC article.
-
Charge state of the globular histone core controls stability of the nucleosome.Biophys J. 2010 Sep 8;99(5):1577-85. doi: 10.1016/j.bpj.2010.06.046. Biophys J. 2010. PMID: 20816070 Free PMC article.
-
Linker histone H1 and H3K56 acetylation are antagonistic regulators of nucleosome dynamics.Nat Commun. 2015 Dec 9;6:10152. doi: 10.1038/ncomms10152. Nat Commun. 2015. PMID: 26648124 Free PMC article.
-
"Direct" and "Indirect" Effects of Histone Modifications: Modulation of Sterical Bulk as a Novel Source of Functionality.Bioessays. 2020 Jan;42(1):e1900136. doi: 10.1002/bies.201900136. Epub 2019 Dec 5. Bioessays. 2020. PMID: 31805213 Review.
-
Histone core modifications regulating nucleosome structure and dynamics.Nat Rev Mol Cell Biol. 2014 Nov;15(11):703-8. doi: 10.1038/nrm3890. Epub 2014 Oct 15. Nat Rev Mol Cell Biol. 2014. PMID: 25315270 Review.
Cited by
-
The cell nucleus. A study in Burgundy.Nucleus. 2019 Dec;10(1):213-217. doi: 10.1080/19491034.2019.1649835. Nucleus. 2019. PMID: 31405317 Free PMC article.
-
Exploring the Potent Anticancer Activity of Essential Oils and Their Bioactive Compounds: Mechanisms and Prospects for Future Cancer Therapy.Pharmaceuticals (Basel). 2023 Jul 31;16(8):1086. doi: 10.3390/ph16081086. Pharmaceuticals (Basel). 2023. PMID: 37631000 Free PMC article. Review.
-
The Vast Complexity of the Epigenetic Landscape during Neurodevelopment: An Open Frame to Understanding Brain Function.Int J Mol Sci. 2018 May 1;19(5):1333. doi: 10.3390/ijms19051333. Int J Mol Sci. 2018. PMID: 29723958 Free PMC article. Review.
-
Cis-regulatory sequences in plants: Their importance, discovery, and future challenges.Plant Cell. 2022 Feb 3;34(2):718-741. doi: 10.1093/plcell/koab281. Plant Cell. 2022. PMID: 34918159 Free PMC article. Review.
-
Semisynthesis of site-specifically succinylated histone reveals that succinylation regulates nucleosome unwrapping rate and DNA accessibility.Nucleic Acids Res. 2020 Sep 25;48(17):9538-9549. doi: 10.1093/nar/gkaa663. Nucleic Acids Res. 2020. PMID: 32766790 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

