Functional architecture of reward learning in mushroom body extrinsic neurons of larval Drosophila
- PMID: 29549237
- PMCID: PMC5856778
- DOI: 10.1038/s41467-018-03130-1
Functional architecture of reward learning in mushroom body extrinsic neurons of larval Drosophila
Abstract
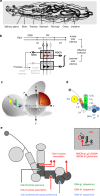
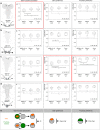
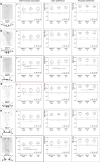
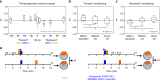
The brain adaptively integrates present sensory input, past experience, and options for future action. The insect mushroom body exemplifies how a central brain structure brings about such integration. Here we use a combination of systematic single-cell labeling, connectomics, transgenic silencing, and activation experiments to study the mushroom body at single-cell resolution, focusing on the behavioral architecture of its input and output neurons (MBINs and MBONs), and of the mushroom body intrinsic APL neuron. Our results reveal the identity and morphology of almost all of these 44 neurons in stage 3 Drosophila larvae. Upon an initial screen, functional analyses focusing on the mushroom body medial lobe uncover sparse and specific functions of its dopaminergic MBINs, its MBONs, and of the GABAergic APL neuron across three behavioral tasks, namely odor preference, taste preference, and associative learning between odor and taste. Our results thus provide a cellular-resolution study case of how brains organize behavior.
Conflict of interest statement
The authors declare no competing financial interests.
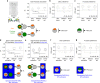
Figures





Similar articles
-
Rewarding Capacity of Optogenetically Activating a Giant GABAergic Central-Brain Interneuron in Larval Drosophila.J Neurosci. 2023 Nov 1;43(44):7393-7428. doi: 10.1523/JNEUROSCI.2310-22.2023. Epub 2023 Sep 21. J Neurosci. 2023. PMID: 37734947 Free PMC article.
-
Four Individually Identified Paired Dopamine Neurons Signal Reward in Larval Drosophila.Curr Biol. 2016 Mar 7;26(5):661-9. doi: 10.1016/j.cub.2016.01.012. Epub 2016 Feb 11. Curr Biol. 2016. PMID: 26877086
-
Metamorphosis of memory circuits in Drosophila reveals a strategy for evolving a larval brain.Elife. 2023 Jan 25;12:e80594. doi: 10.7554/eLife.80594. Elife. 2023. PMID: 36695420 Free PMC article.
-
Odor-taste learning in Drosophila larvae.J Insect Physiol. 2018 Apr;106(Pt 1):47-54. doi: 10.1016/j.jinsphys.2017.08.004. Epub 2017 Aug 18. J Insect Physiol. 2018. PMID: 28823531 Review.
-
Connectomics and function of a memory network: the mushroom body of larval Drosophila.Curr Opin Neurobiol. 2019 Feb;54:146-154. doi: 10.1016/j.conb.2018.10.007. Epub 2018 Oct 24. Curr Opin Neurobiol. 2019. PMID: 30368037 Review.
Cited by
-
Pruning deficits of the developing Drosophila mushroom body result in mild impairment in associative odour learning and cause hyperactivity.Open Biol. 2022 Sep;12(9):220096. doi: 10.1098/rsob.220096. Epub 2022 Sep 21. Open Biol. 2022. PMID: 36128716 Free PMC article.
-
Modulations of microbehaviour by associative memory strength in Drosophila larvae.PLoS One. 2019 Oct 21;14(10):e0224154. doi: 10.1371/journal.pone.0224154. eCollection 2019. PLoS One. 2019. PMID: 31634372 Free PMC article.
-
Linking neural circuits to the mechanics of animal behavior in Drosophila larval locomotion.Front Neural Circuits. 2023 Aug 17;17:1175899. doi: 10.3389/fncir.2023.1175899. eCollection 2023. Front Neural Circuits. 2023. PMID: 37711343 Free PMC article. Review.
-
High-resolution analysis of individual Drosophila melanogaster larvae uncovers individual variability in locomotion and its neurogenetic modulation.Open Biol. 2023 Apr;13(4):220308. doi: 10.1098/rsob.220308. Epub 2023 Apr 19. Open Biol. 2023. PMID: 37072034 Free PMC article.
-
Serotonin receptor 5-HT7 in Drosophila mushroom body neurons mediates larval appetitive olfactory learning.Sci Rep. 2020 Dec 4;10(1):21267. doi: 10.1038/s41598-020-77910-5. Sci Rep. 2020. PMID: 33277559 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

