Monoclonal antibody targeting the β-barrel assembly machine of Escherichia coli is bactericidal
- PMID: 29555747
- PMCID: PMC5889671
- DOI: 10.1073/pnas.1800043115
Monoclonal antibody targeting the β-barrel assembly machine of Escherichia coli is bactericidal
Abstract
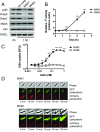
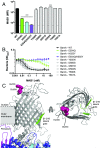
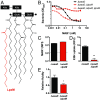
The folding and insertion of integral β-barrel membrane proteins into the outer membrane of Gram-negative bacteria is required for viability and bacterial pathogenesis. Unfortunately, the lack of selective and potent modulators to dissect β-barrel folding in vivo has hampered our understanding of this fundamental biological process. Here, we characterize a monoclonal antibody that selectively inhibits an essential component of the Escherichia coli β-barrel assembly machine, BamA. In the absence of complement or other immune factors, the unmodified antibody MAB1 demonstrates bactericidal activity against an E. coli strain with truncated LPS. Direct binding of MAB1 to an extracellular BamA epitope inhibits its β-barrel folding activity, induces periplasmic stress, disrupts outer membrane integrity, and kills bacteria. Notably, resistance to MAB1-mediated killing reveals a link between outer membrane fluidity and protein folding by BamA in vivo, underscoring the utility of this antibody for studying β-barrel membrane protein folding within a living cell. Identification of this BamA antagonist highlights the potential for new mechanisms of antibiotics to inhibit Gram-negative bacterial growth by targeting extracellular epitopes.
Keywords: BamA; Gram-negative bacteria; LPS; membrane protein folding; β-barrel protein.
Copyright © 2018 the Author(s). Published by PNAS.
Conflict of interest statement
Conflict of interest statement: All authors are employees of Genentech, a member of the Roche Group, and are shareholders in Roche. Study was supported by internal Genentech funds, and the funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Figures
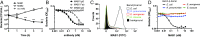
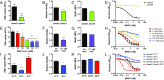
Comment in
-
Directly Bactericidal Anti-Escherichia coli Antibody.Trends Microbiol. 2018 Aug;26(8):642-644. doi: 10.1016/j.tim.2018.05.016. Epub 2018 Jun 11. Trends Microbiol. 2018. PMID: 29903418
Similar articles
-
The Escherichia coli β-Barrel Assembly Machinery Is Sensitized to Perturbations under High Membrane Fluidity.J Bacteriol. 2018 Dec 7;201(1):e00517-18. doi: 10.1128/JB.00517-18. Print 2019 Jan 1. J Bacteriol. 2018. PMID: 30322857 Free PMC article.
-
A small-molecule inhibitor of BamA impervious to efflux and the outer membrane permeability barrier.Proc Natl Acad Sci U S A. 2019 Oct 22;116(43):21748-21757. doi: 10.1073/pnas.1912345116. Epub 2019 Oct 7. Proc Natl Acad Sci U S A. 2019. PMID: 31591200 Free PMC article.
-
A Peptidomimetic Antibiotic Targets Outer Membrane Proteins and Disrupts Selectively the Outer Membrane in Escherichia coli.J Biol Chem. 2016 Jan 22;291(4):1921-1932. doi: 10.1074/jbc.M115.691725. Epub 2015 Dec 1. J Biol Chem. 2016. PMID: 26627837 Free PMC article.
-
Augmenting β-augmentation: structural basis of how BamB binds BamA and may support folding of outer membrane proteins.J Mol Biol. 2011 Mar 11;406(5):659-66. doi: 10.1016/j.jmb.2011.01.002. Epub 2011 Jan 12. J Mol Biol. 2011. PMID: 21236263 Review.
-
The β-Barrel Assembly Machinery Complex.Methods Mol Biol. 2015;1329:1-16. doi: 10.1007/978-1-4939-2871-2_1. Methods Mol Biol. 2015. PMID: 26427672 Review.
Cited by
-
Alternative therapeutic strategies to treat antibiotic-resistant pathogens.Nat Rev Microbiol. 2024 May;22(5):262-275. doi: 10.1038/s41579-023-00993-0. Epub 2023 Dec 11. Nat Rev Microbiol. 2024. PMID: 38082064 Review.
-
Darobactin B Stabilises a Lateral-Closed Conformation of the BAM Complex in E. coli Cells.Angew Chem Weinheim Bergstr Ger. 2023 Aug 21;135(34):e202218783. doi: 10.1002/ange.202218783. Epub 2023 May 31. Angew Chem Weinheim Bergstr Ger. 2023. PMID: 38515502 Free PMC article.
-
β-Barrel Assembly Machinery (BAM) Complex as Novel Antibacterial Drug Target.Molecules. 2023 Apr 27;28(9):3758. doi: 10.3390/molecules28093758. Molecules. 2023. PMID: 37175168 Free PMC article. Review.
-
Immobile lipopolysaccharides and outer membrane proteins differentially segregate in growing Escherichia coli.Proc Natl Acad Sci U S A. 2025 Mar 11;122(10):e2414725122. doi: 10.1073/pnas.2414725122. Epub 2025 Mar 3. Proc Natl Acad Sci U S A. 2025. PMID: 40030021 Free PMC article.
-
Engineered peptide barcodes for in-depth analyses of binding protein libraries.Nat Methods. 2019 May;16(5):421-428. doi: 10.1038/s41592-019-0389-8. Epub 2019 Apr 22. Nat Methods. 2019. PMID: 31011184 Free PMC article.
References
-
- Kamio Y, Nikaido H. Outer membrane of Salmonella typhimurium: Accessibility of phospholipid head groups to phospholipase c and cyanogen bromide activated dextran in the external medium. Biochemistry. 1976;15:2561–2570. - PubMed
-
- Kleinschmidt JH. Folding of β-barrel membrane proteins in lipid bilayers—Unassisted and assisted folding and insertion. Biochim Biophys Acta. 2015;1848:1927–1943. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

