The Colibactin Genotoxin Generates DNA Interstrand Cross-Links in Infected Cells
- PMID: 29559578
- PMCID: PMC5874909
- DOI: 10.1128/mBio.02393-17
The Colibactin Genotoxin Generates DNA Interstrand Cross-Links in Infected Cells
Abstract
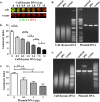
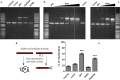
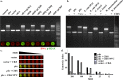
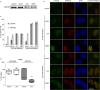
Colibactins are hybrid polyketide-nonribosomal peptides produced by Escherichia coli, Klebsiella pneumoniae, and other Enterobacteriaceae harboring the pks genomic island. These genotoxic metabolites are produced by pks-encoded peptide-polyketide synthases as inactive prodrugs called precolibactins, which are then converted to colibactins by deacylation for DNA-damaging effects. Colibactins are bona fide virulence factors and are suspected of promoting colorectal carcinogenesis when produced by intestinal E. coli Natural active colibactins have not been isolated, and how they induce DNA damage in the eukaryotic host cell is poorly characterized. Here, we show that DNA strands are cross-linked covalently when exposed to enterobacteria producing colibactins. DNA cross-linking is abrogated in a clbP mutant unable to deacetylate precolibactins or by adding the colibactin self-resistance protein ClbS, confirming the involvement of the mature forms of colibactins. A similar DNA-damaging mechanism is observed in cellulo, where interstrand cross-links are detected in the genomic DNA of cultured human cells exposed to colibactin-producing bacteria. The intoxicated cells exhibit replication stress, activation of ataxia-telangiectasia and Rad3-related kinase (ATR), and recruitment of the DNA cross-link repair Fanconi anemia protein D2 (FANCD2) protein. In contrast, inhibition of ATR or knockdown of FANCD2 reduces the survival of cells exposed to colibactin-producing bacteria. These findings demonstrate that DNA interstrand cross-linking is the critical mechanism of colibactin-induced DNA damage in infected cells.IMPORTANCE Colorectal cancer is the third-most-common cause of cancer death. In addition to known risk factors such as high-fat diets and alcohol consumption, genotoxic intestinal Escherichia coli bacteria producing colibactin are proposed to play a role in colon cancer development. Here, by using transient infections with genotoxic E. coli, we showed that colibactins directly generate DNA cross-links in cellulo Such lesions are converted into double-strand breaks during the repair response. DNA cross-links, akin to those induced by metabolites of alcohol and high-fat diets and by widely used anticancer drugs, are both severely mutagenic and profoundly cytotoxic lesions. This finding of a direct induction of DNA cross-links by a bacterium should facilitate delineating the role of E. coli in colon cancer and engineering new anticancer agents.
Keywords: DNA cross-linking agents; DNA damage; DNA damage checkpoints; Escherichia coli; Escherichia toxins; genotoxicity.
Copyright © 2018 Bossuet-Greif et al.
Figures

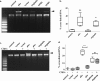
References
-
- Putze J, Hennequin C, Nougayrède JP, Zhang W, Homburg S, Karch H, Bringer MA, Fayolle C, Carniel E, Rabsch W, Oelschlaeger TA, Oswald E, Forestier C, Hacker J, Dobrindt U. 2009. Genetic structure and distribution of the colibactin genomic island among members of the family Enterobacteriaceae. Infect Immun 77:4696–4703. doi:10.1128/IAI.00522-09. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

