SuPReMe: a rapid reverse genetics method to generate clonal populations of recombinant RNA viruses
- PMID: 29559627
- PMCID: PMC5861084
- DOI: 10.1038/s41426-018-0040-2
SuPReMe: a rapid reverse genetics method to generate clonal populations of recombinant RNA viruses
Abstract
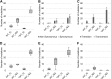
Reverse genetics systems enable the manipulation of viral genomes and are proving to be essential for studying RNA viruses. Methods for generating clonal virus populations are particularly useful for studying the impact of genomic modifications on viral properties. Here, by exploiting a chikungunya virus model, we compare viral populations and their replicative fitness when generated using either the rapid and user-friendly PCR-based ISA (Infectious Subgenomic Amplicons) method or classical infectious clone technology. As anticipated, the ISA method resulted in greater genetic diversity of the viral populations, but no significant difference in viral fitness in vitro was observed. On the basis of these results, a new ISA-derived reverse genetics procedure was developed. This method, designated 'SuPReMe' (Subgenomic Plasmids Recombination Method), in which digested plasmids containing subgenomic DNA fragments were directly transfected into permissive cells, retains the following major advantages of the ISA method: it is rapid, flexible and does not require the cloning of complete genomes. Moreover, SuPReMe has been shown to produce virus populations with genetic diversity and replicative fitness similar to those obtained using conventional infectious clone technology. SuPReMe, therefore, represents an effective and promising option for the rapid generation of clonal recombinant populations of single-stranded positive-sense RNA viruses.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


Similar articles
-
Reverse Genetics of RNA Viruses: ISA-Based Approach to Control Viral Population Diversity without Modifying Virus Phenotype.Viruses. 2019 Jul 20;11(7):666. doi: 10.3390/v11070666. Viruses. 2019. PMID: 31330809 Free PMC article.
-
Comparison of chikungunya viruses generated using infectious clone or the Infectious Subgenomic Amplicons (ISA) method in Aedes mosquitoes.PLoS One. 2018 Jun 28;13(6):e0199494. doi: 10.1371/journal.pone.0199494. eCollection 2018. PLoS One. 2018. PMID: 29953474 Free PMC article.
-
In vivo rescue of arboviruses directly from subgenomic DNA fragments.Emerg Microbes Infect. 2024 Dec;13(1):2356140. doi: 10.1080/22221751.2024.2356140. Epub 2024 May 26. Emerg Microbes Infect. 2024. PMID: 38742328 Free PMC article.
-
Highs and Lows in Calicivirus Reverse Genetics.Viruses. 2024 May 28;16(6):866. doi: 10.3390/v16060866. Viruses. 2024. PMID: 38932159 Free PMC article. Review.
-
Reverse genetics system for human rotaviruses.Microbiol Immunol. 2020 Jun;64(6):401-406. doi: 10.1111/1348-0421.12795. Epub 2020 May 2. Microbiol Immunol. 2020. PMID: 32293752 Review.
Cited by
-
A combination of two resistance mechanisms is critical for tick-borne encephalitis virus escape from a broadly neutralizing human antibody.Cell Rep. 2023 Sep 26;42(9):113149. doi: 10.1016/j.celrep.2023.113149. Epub 2023 Sep 19. Cell Rep. 2023. PMID: 37715951 Free PMC article.
-
Development and Characterization of a Genetically Stable Infectious Clone for a Genotype I Isolate of Dengue Virus Serotype 1.Viruses. 2022 Sep 18;14(9):2073. doi: 10.3390/v14092073. Viruses. 2022. PMID: 36146879 Free PMC article.
-
Rapid cloning-free mutagenesis of new SARS-CoV-2 variants using a novel reverse genetics platform.Elife. 2023 Nov 21;12:RP89035. doi: 10.7554/eLife.89035. Elife. 2023. PMID: 37988285 Free PMC article.
-
Reverse Genetics of RNA Viruses: ISA-Based Approach to Control Viral Population Diversity without Modifying Virus Phenotype.Viruses. 2019 Jul 20;11(7):666. doi: 10.3390/v11070666. Viruses. 2019. PMID: 31330809 Free PMC article.
-
An E460D Substitution in the NS5 Protein of Tick-Borne Encephalitis Virus Confers Resistance to the Inhibitor Galidesivir (BCX4430) and Also Attenuates the Virus for Mice.J Virol. 2019 Jul 30;93(16):e00367-19. doi: 10.1128/JVI.00367-19. Print 2019 Aug 15. J Virol. 2019. PMID: 31142664 Free PMC article.
References
-
- Neumann G, Kawaoka Y. Reverse genetics systems for the generation of segmented negative-sense RNA viruses entirely from cloned cDNA. Curr. Top. Microbiol Immunol. 2004;283:43–60. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
