Measuring and Estimating the Effect Sizes of Copy Number Variants on General Intelligence in Community-Based Samples
- PMID: 29562078
- PMCID: PMC5875373
- DOI: 10.1001/jamapsychiatry.2018.0039
Measuring and Estimating the Effect Sizes of Copy Number Variants on General Intelligence in Community-Based Samples
Abstract
Importance;: Copy number variants (CNVs) classified as pathogenic are identified in 10% to 15% of patients referred for neurodevelopmental disorders. However, their effect sizes on cognitive traits measured as a continuum remain mostly unknown because most of them are too rare to be studied individually using association studies.
Objective: To measure and estimate the effect sizes of recurrent and nonrecurrent CNVs on IQ.
Design, setting, and participants: This study identified all CNVs that were 50 kilobases (kb) or larger in 2 general population cohorts (the IMAGEN project and the Saguenay Youth Study) with measures of IQ. Linear regressions, including functional annotations of genes included in CNVs, were used to identify features to explain their association with IQ. Validation was performed using intraclass correlation that compared IQ estimated by the model with empirical data.
Main outcomes and measures: Performance IQ (PIQ), verbal IQ (VIQ), and frequency of de novo CNV events.
Results: The study included 2090 European adolescents from the IMAGEN study and 1983 children and parents from the Saguenay Youth Study. Of these, genotyping was performed on 1804 individuals from IMAGEN and 977 adolescents, 445 mothers, and 448 fathers (484 families) from the Saguenay Youth Study. We observed 4928 autosomal CNVs larger than 50 kb across both cohorts. For rare deletions, size, number of genes, and exons affect IQ, and each deleted gene is associated with a mean (SE) decrease in PIQ of 0.67 (0.19) points (P = 6 × 10-4); this is not so for rare duplications and frequent CNVs. Among 10 functional annotations, haploinsufficiency scores best explain the association of any deletions with PIQ with a mean (SE) decrease of 2.74 (0.68) points per unit of the probability of being loss-of-function intolerant (P = 8 × 10-5). Results are consistent across cohorts and unaffected by sensitivity analyses removing pathogenic CNVs. There is a 0.75 concordance (95% CI, 0.39-0.91) between the effect size on IQ estimated by our model and IQ loss calculated in previous studies of 15 recurrent CNVs. There is a close association between effect size on IQ and the frequency at which deletions occur de novo (odds ratio, 0.86; 95% CI, 0.84-0.87; P = 2.7 × 10-88). There is a 0.76 concordance (95% CI, 0.41-0.91) between de novo frequency estimated by the model and calculated using data from the DECIPHER database.
Conclusions and relevance: Models trained on nonpathogenic deletions in the general population reliably estimate the effect size of pathogenic deletions and suggest omnigenic associations of haploinsufficiency with IQ. This represents a new framework to study variants too rare to perform individual association studies and can help estimate the cognitive effect of undocumented deletions in the neurodevelopmental clinic.
Conflict of interest statement
Figures

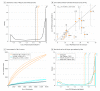
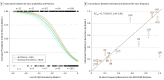
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

