RNA cytosine methylation and methyltransferases mediate chromatin organization and 5-azacytidine response and resistance in leukaemia
- PMID: 29563491
- PMCID: PMC5862959
- DOI: 10.1038/s41467-018-03513-4
RNA cytosine methylation and methyltransferases mediate chromatin organization and 5-azacytidine response and resistance in leukaemia
Erratum in
-
Author Correction: RNA cytosine methylation and methyltransferases mediate chromatin organization and 5-azacytidine response and resistance in leukaemia.Nat Commun. 2018 Jun 6;9(1):2286. doi: 10.1038/s41467-018-04518-9. Nat Commun. 2018. PMID: 29875356 Free PMC article.
Abstract
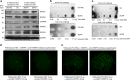
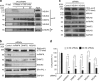
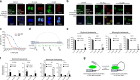
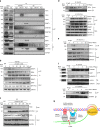
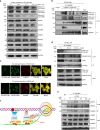
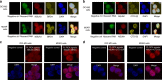
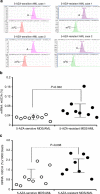
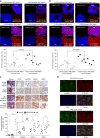
The roles of RNA 5-methylcytosine (RNA:m5C) and RNA:m5C methyltransferases (RCMTs) in lineage-associated chromatin organization and drug response/resistance are unclear. Here we demonstrate that the RCMTs, namely NSUN3 and DNMT2, directly bind hnRNPK, a conserved RNA-binding protein. hnRNPK interacts with the lineage-determining transcription factors (TFs), GATA1 and SPI1/PU.1, and with CDK9/P-TEFb to recruit RNA-polymerase-II at nascent RNA, leading to formation of 5-Azacitidine (5-AZA)-sensitive chromatin structure. In contrast, NSUN1 binds BRD4 and RNA-polymerase-II to form an active chromatin structure that is insensitive to 5-AZA, but hypersensitive to the BRD4 inhibitor JQ1 and to the downregulation of NSUN1 by siRNAs. Both 5-AZA-resistant leukaemia cell lines and clinically 5-AZA-resistant myelodysplastic syndrome and acute myeloid leukaemia specimens have a significant increase in RNA:m5C and NSUN1-/BRD4-associated active chromatin. This study reveals novel RNA:m5C/RCMT-mediated chromatin structures that modulate 5-AZA response/resistance in leukaemia cells, and hence provides a new insight into treatment of leukaemia.
Conflict of interest statement
The authors declare no competing interests.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

