Selective Enrichment and Direct Analysis of Protein S-Palmitoylation Sites
- PMID: 29575903
- PMCID: PMC6104640
- DOI: 10.1021/acs.jproteome.8b00002
Selective Enrichment and Direct Analysis of Protein S-Palmitoylation Sites
Abstract
S-Fatty-acylation is the covalent attachment of long chain fatty acids, predominately palmitate (C16:0, S-palmitoylation), to cysteine (Cys) residues via a thioester linkage on proteins. This post-translational and reversible lipid modification regulates protein function and localization in eukaryotes and is important in mammalian physiology and human diseases. While chemical labeling methods have improved the detection and enrichment of S-fatty-acylated proteins, mapping sites of modification and characterizing the endogenously attached fatty acids are still challenging. Here, we describe the integration and optimization of fatty acid chemical reporter labeling with hydroxylamine-mediated enrichment of S-fatty-acylated proteins and direct tagging of modified Cys residues to selectively map lipid modification sites. This afforded improved enrichment and direct identification of many protein S-fatty-acylation sites compared to previously described methods. Notably, we directly identified the S-fatty-acylation sites of IFITM3, an important interferon-stimulated inhibitor of virus entry, and we further demonstrated that the highly conserved Cys residues are primarily modified by palmitic acid. The methods described here should facilitate the direct analysis of protein S-fatty-acylation sites and their endogenously attached fatty acids in diverse cell types and activation states important for mammalian physiology and diseases.
Keywords: S-palmitoylation; affinity enrichment; fatty acylation; mass spectrometry-based proteomics; posttranslational modification; site identification.
Figures



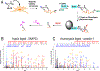

Similar articles
-
Analysis of Protein Cysteine Acylation Using a Modified Suspension Trap (Acyl-Trap).J Proteome Res. 2024 Aug 2;23(8):3716-3725. doi: 10.1021/acs.jproteome.4c00225. Epub 2024 Jul 15. J Proteome Res. 2024. PMID: 39008777 Free PMC article.
-
Chemical Proteomic Analysis of S-Fatty Acylated Proteins and Their Modification Sites.Methods Mol Biol. 2019;2009:45-57. doi: 10.1007/978-1-4939-9532-5_4. Methods Mol Biol. 2019. PMID: 31152394
-
Mass-tag labeling reveals site-specific and endogenous levels of protein S-fatty acylation.Proc Natl Acad Sci U S A. 2016 Apr 19;113(16):4302-7. doi: 10.1073/pnas.1602244113. Epub 2016 Apr 4. Proc Natl Acad Sci U S A. 2016. PMID: 27044110 Free PMC article.
-
Differential S-Acylation of Enveloped Viruses.Protein Pept Lett. 2019;26(8):588-600. doi: 10.2174/0929866526666190603082521. Protein Pept Lett. 2019. PMID: 31161979 Review.
-
The physiology of protein S-acylation.Physiol Rev. 2015 Apr;95(2):341-76. doi: 10.1152/physrev.00032.2014. Physiol Rev. 2015. PMID: 25834228 Free PMC article. Review.
Cited by
-
Protein S-Acyl Transferase GhPAT27 Was Associated with Verticillium wilt Resistance in Cotton.Plants (Basel). 2022 Oct 18;11(20):2758. doi: 10.3390/plants11202758. Plants (Basel). 2022. PMID: 36297782 Free PMC article.
-
Palmitoylated importin α regulates mitotic spindle orientation through interaction with NuMA.EMBO Rep. 2025 Jul;26(13):3280-3304. doi: 10.1038/s44319-025-00484-8. Epub 2025 May 27. EMBO Rep. 2025. PMID: 40425783 Free PMC article.
-
Discovery and Characterization of IFITM S-Palmitoylation.Viruses. 2023 Nov 28;15(12):2329. doi: 10.3390/v15122329. Viruses. 2023. PMID: 38140570 Free PMC article. Review.
-
Site-Specific Lipidation Enhances IFITM3 Membrane Interactions and Antiviral Activity.ACS Chem Biol. 2021 May 21;16(5):844-856. doi: 10.1021/acschembio.1c00013. Epub 2021 Apr 22. ACS Chem Biol. 2021. PMID: 33887136 Free PMC article.
-
Analysis of Protein Cysteine Acylation Using a Modified Suspension Trap (Acyl-Trap).J Proteome Res. 2024 Aug 2;23(8):3716-3725. doi: 10.1021/acs.jproteome.4c00225. Epub 2024 Jul 15. J Proteome Res. 2024. PMID: 39008777 Free PMC article.
References
-
- Lanyon-Hogg T; Faronato M; Serwa RA; Tate EW Dynamic Protein Acylation: New Substrates, Mechanisms, and Drug Targets. Trends Biochem. Sci 2017, 42 (7), 566–581. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

