Non-Pleiotropic Coupling of Daily and Seasonal Temporal Isolation in the European Corn Borer
- PMID: 29587435
- PMCID: PMC5924522
- DOI: 10.3390/genes9040180
Non-Pleiotropic Coupling of Daily and Seasonal Temporal Isolation in the European Corn Borer
Abstract
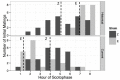
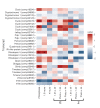
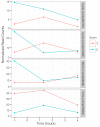
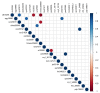
Speciation often involves the coupling of multiple isolating barriers to produce reproductive isolation, but how coupling is generated among different premating barriers is unknown. We measure the degree of coupling between the daily mating time and seasonal mating time between strains of European corn borer (Ostrinia nubilalis) and evaluate the hypothesis that the coupling of different forms of allochrony is due to a shared genetic architecture, involving genes with pleiotropic effects on both timing phenotypes. We measure differences in gene expression at peak mating times and compare these genes to previously identified candidates that are associated with changes in seasonal mating time between the corn borer strains. We find that the E strain, which mates earlier in the season, also mates 2.7 h earlier in the night than the Z strain. Earlier daily mating is correlated with the differences in expression of the circadian clock genes cycle, slimb, and vrille. However, different circadian clock genes associate with daily and seasonal timing, suggesting that the coupling of timing traits is maintained by natural selection rather than pleiotropy. Juvenile hormone gene expression was associated with both types of timing, suggesting that circadian genes activate common downstream modules that may impose constraint on future evolution of these traits.
Keywords: allochronic isolation; circadian clock; gene expression; sexual behavior; speciation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Coyne J.A., Orr H.A. Speciation. Volume 37 Sinauer Associates; Sunderland, MA, USA: 2004.
LinkOut - more resources
Full Text Sources
Other Literature Sources

