Nascent DNA methylome mapping reveals inheritance of hemimethylation at CTCF/cohesin sites
- PMID: 29590048
- PMCID: PMC6359960
- DOI: 10.1126/science.aan5480
Nascent DNA methylome mapping reveals inheritance of hemimethylation at CTCF/cohesin sites
Abstract
The faithful inheritance of the epigenome is critical for cells to maintain gene expression programs and cellular identity across cell divisions. We mapped strand-specific DNA methylation after replication forks and show maintenance of the vast majority of the DNA methylome within 20 minutes of replication and inheritance of some hemimethylated CpG dinucleotides (hemiCpGs). Mapping the nascent DNA methylome targeted by each of the three DNA methyltransferases (DNMTs) reveals interactions between DNMTs and substrate daughter cytosines en route to maintenance methylation or hemimethylation. Finally, we show the inheritance of hemiCpGs at short regions flanking CCCTC-binding factor (CTCF)/cohesin binding sites in pluripotent cells. Elimination of hemimethylation causes reduced frequency of chromatin interactions emanating from these sites, suggesting a role for hemimethylation as a stable epigenetic mark regulating CTCF-mediated chromatin interactions.
Copyright © 2018 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Figures

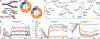
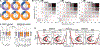
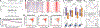
Comment in
-
Hemimethylation: DNA's lasting odd couple.Science. 2018 Mar 9;359(6380):1102-1103. doi: 10.1126/science.aat0789. Science. 2018. PMID: 29590029 No abstract available.
References
-
- Holliday R, Pugh JE, DNA modification mechanisms and gene activity during development. Science 187, 226–232 (1975). - PubMed
-
- Chuang LS et al., Human DNA-(cytosine-5) methyltransferase-PCNA complex as a target for p21WAFl. Science 277, 1996–2000 (1997). - PubMed
-
- Bostick M et al., UHRF1 plays a role in maintaining DNA methylation in mammalian cells. Science 317, 1760–1764 (2007). - PubMed
-
- Sharif J et al., The SRA protein Np95 mediates epigenetic inheritance by recruiting Dnmtl to methylated DNA. Nature 450, 908–912 (2007). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

