Within-Host Variations of Human Papillomavirus Reveal APOBEC Signature Mutagenesis in the Viral Genome
- PMID: 29593040
- PMCID: PMC5974501
- DOI: 10.1128/JVI.00017-18
Within-Host Variations of Human Papillomavirus Reveal APOBEC Signature Mutagenesis in the Viral Genome
Abstract
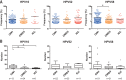
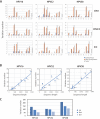
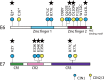
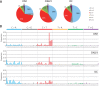
Persistent infection with oncogenic human papillomaviruses (HPVs) causes cervical cancer, accompanied by the accumulation of somatic mutations into the host genome. There are concomitant genetic changes in the HPV genome during viral infection; however, their relevance to cervical carcinogenesis is poorly understood. Here, we explored within-host genetic diversity of HPV by performing deep-sequencing analyses of viral whole-genome sequences in clinical specimens. The whole genomes of HPV types 16, 52, and 58 were amplified by type-specific PCR from total cellular DNA of cervical exfoliated cells collected from patients with cervical intraepithelial neoplasia (CIN) and invasive cervical cancer (ICC) and were deep sequenced. After constructing a reference viral genome sequence for each specimen, nucleotide positions showing changes with >0.5% frequencies compared to the reference sequence were determined for individual samples. In total, 1,052 positions of nucleotide variations were detected in HPV genomes from 151 samples (CIN1, n = 56; CIN2/3, n = 68; ICC, n = 27), with various numbers per sample. Overall, C-to-T and C-to-A substitutions were the dominant changes observed across all histological grades. While C-to-T transitions were predominantly detected in CIN1, their prevalence was decreased in CIN2/3 and fell below that of C-to-A transversions in ICC. Analysis of the trinucleotide context encompassing substituted bases revealed that TpCpN, a preferred target sequence for cellular APOBEC cytosine deaminases, was a primary site for C-to-T substitutions in the HPV genome. These results strongly imply that the APOBEC proteins are drivers of HPV genome mutation, particularly in CIN1 lesions.IMPORTANCE HPVs exhibit surprisingly high levels of genetic diversity, including a large repertoire of minor genomic variants in each viral genotype. Here, by conducting deep-sequencing analyses, we show for the first time a comprehensive snapshot of the within-host genetic diversity of high-risk HPVs during cervical carcinogenesis. Quasispecies harboring minor nucleotide variations in viral whole-genome sequences were extensively observed across different grades of CIN and cervical cancer. Among the within-host variations, C-to-T transitions, a characteristic change mediated by cellular APOBEC cytosine deaminases, were predominantly detected throughout the whole viral genome, most strikingly in low-grade CIN lesions. The results strongly suggest that within-host variations of the HPV genome are primarily generated through the interaction with host cell DNA-editing enzymes and that such within-host variability is an evolutionary source of the genetic diversity of HPVs.
Keywords: APOBEC; genetic diversity; human papillomavirus; next-generation sequencing; quasispecies.
Copyright © 2018 American Society for Microbiology.
Figures

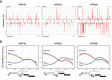
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

