Comparative Genomics and Biosynthetic Potential Analysis of Two Lichen-Isolated Amycolatopsis Strains
- PMID: 29593664
- PMCID: PMC5859366
- DOI: 10.3389/fmicb.2018.00369
Comparative Genomics and Biosynthetic Potential Analysis of Two Lichen-Isolated Amycolatopsis Strains
Abstract
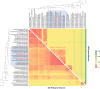
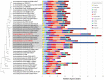
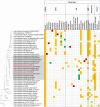
Actinomycetes have been extensively exploited as one of the most prolific secondary metabolite-producer sources and continue to be in the focus of interest in the constant search of novel bioactive compounds. The availability of less expensive next generation genome sequencing techniques has not only confirmed the extraordinary richness and broad distribution of silent natural product biosynthetic gene clusters among these bacterial genomes, but also has allowed the incorporation of genomics in bacterial taxonomy and systematics. As part of our efforts to isolate novel strains from unique environments, we explored lichen-associated microbial communities as unique assemblages to be studied as potential sources of novel bioactive natural products with application in biotechnology and drug discovery. In this work, we have studied the whole genome sequences of two new Amycolatopsis strains (CA-126428 and CA-128772) isolated from tropical lichens, and performed a comparative genomic analysis with 41 publicly available Amycolatopsis genomes. This work has not only permitted to infer and discuss their taxonomic position on the basis of the different phylogenetic approaches used, but has also allowed to assess the richness and uniqueness of the biosynthetic pathways associated to primary and secondary metabolism, and to provide a first insight on the potential role of these bacteria in the lichen-associated microbial community.
Keywords: Amycolatopsis; biosynthetic gene clusters; phylogeny; secondary metabolites; whole genome sequence.
Figures



Similar articles
-
Comparative genomics reveals phylogenetic distribution patterns of secondary metabolites in Amycolatopsis species.BMC Genomics. 2018 Jun 1;19(1):426. doi: 10.1186/s12864-018-4809-4. BMC Genomics. 2018. PMID: 29859036 Free PMC article.
-
A Genomic Survey of the Natural Product Biosynthetic Potential of Actinomycetes Isolated from New Zealand Lichens.mSystems. 2023 Apr 27;8(2):e0103022. doi: 10.1128/msystems.01030-22. Epub 2023 Feb 7. mSystems. 2023. PMID: 36749048 Free PMC article.
-
Biosynthetic Gene Content of the 'Perfume Lichens' Evernia prunastri and Pseudevernia furfuracea.Molecules. 2019 Jan 8;24(1):203. doi: 10.3390/molecules24010203. Molecules. 2019. PMID: 30626017 Free PMC article.
-
Secondary Metabolites of the Genus Amycolatopsis: Structures, Bioactivities and Biosynthesis.Molecules. 2021 Mar 26;26(7):1884. doi: 10.3390/molecules26071884. Molecules. 2021. PMID: 33810439 Free PMC article. Review.
-
Secondary metabolism in the lichen symbiosis.Chem Soc Rev. 2018 Mar 5;47(5):1730-1760. doi: 10.1039/c7cs00431a. Chem Soc Rev. 2018. PMID: 29094129 Review.
Cited by
-
Revisiting the Taxonomic Status of the Biomedically and Industrially Important Genus Amycolatopsis, Using a Phylogenomic Approach.Front Microbiol. 2018 Sep 27;9:2281. doi: 10.3389/fmicb.2018.02281. eCollection 2018. Front Microbiol. 2018. PMID: 30319584 Free PMC article.
-
The Degradative Capabilities of New Amycolatopsis Isolates on Polylactic Acid.Microorganisms. 2019 Nov 20;7(12):590. doi: 10.3390/microorganisms7120590. Microorganisms. 2019. PMID: 31757055 Free PMC article.
-
Chronicle of Research into Lichen-Associated Bacteria.Microorganisms. 2022 Oct 26;10(11):2111. doi: 10.3390/microorganisms10112111. Microorganisms. 2022. PMID: 36363703 Free PMC article. Review.
-
Amycolatopsis camponoti sp. nov., new tetracenomycin-producing actinomycete isolated from carpenter ant Camponotus vagus.Antonie Van Leeuwenhoek. 2022 Apr;115(4):533-544. doi: 10.1007/s10482-022-01716-w. Epub 2022 Feb 26. Antonie Van Leeuwenhoek. 2022. PMID: 35218449 Free PMC article.
-
Mining Actinomycetes for Novel Antibiotics in the Omics Era: Are We Ready to Exploit This New Paradigm?Antibiotics (Basel). 2018 Sep 25;7(4):85. doi: 10.3390/antibiotics7040085. Antibiotics (Basel). 2018. PMID: 30257490 Free PMC article. Review.
References
-
- Adamek M., Spohn M., Stegmann E., Ziemert N. (2017). Mining bacterial genomes for secondary metabolite gene clusters, in Antibiotics, Methods in Molecular Biology, ed Sass P. (New York, NY: Humana Press; ), 23–47. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

