Flavin-Based Electron Bifurcation, Ferredoxin, Flavodoxin, and Anaerobic Respiration With Protons (Ech) or NAD+ (Rnf) as Electron Acceptors: A Historical Review
- PMID: 29593673
- PMCID: PMC5861303
- DOI: 10.3389/fmicb.2018.00401
Flavin-Based Electron Bifurcation, Ferredoxin, Flavodoxin, and Anaerobic Respiration With Protons (Ech) or NAD+ (Rnf) as Electron Acceptors: A Historical Review
Abstract
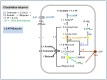
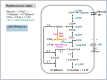
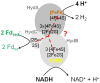
Flavin-based electron bifurcation is a newly discovered mechanism, by which a hydride electron pair from NAD(P)H, coenzyme F420H2, H2, or formate is split by flavoproteins into one-electron with a more negative reduction potential and one with a more positive reduction potential than that of the electron pair. Via this mechanism microorganisms generate low- potential electrons for the reduction of ferredoxins (Fd) and flavodoxins (Fld). The first example was described in 2008 when it was found that the butyryl-CoA dehydrogenase-electron-transferring flavoprotein complex (Bcd-EtfAB) of Clostridium kluyveri couples the endergonic reduction of ferredoxin (E0' = -420 mV) with NADH (-320 mV) to the exergonic reduction of crotonyl-CoA to butyryl-CoA (-10 mV) with NADH. The discovery was followed by the finding of an electron-bifurcating Fd- and NAD-dependent [FeFe]-hydrogenase (HydABC) in Thermotoga maritima (2009), Fd-dependent transhydrogenase (NfnAB) in various bacteria and archaea (2010), Fd- and H2-dependent heterodisulfide reductase (MvhADG-HdrABC) in methanogenic archaea (2011), Fd- and NADH-dependent caffeyl-CoA reductase (CarCDE) in Acetobacterium woodii (2013), Fd- and NAD-dependent formate dehydrogenase (HylABC-FdhF2) in Clostridium acidi-urici (2013), Fd- and NADP-dependent [FeFe]-hydrogenase (HytA-E) in Clostridium autoethanogrenum (2013), Fd(?)- and NADH-dependent methylene-tetrahydrofolate reductase (MetFV-HdrABC-MvhD) in Moorella thermoacetica (2014), Fd- and NAD-dependent lactate dehydrogenase (LctBCD) in A. woodii (2015), Fd- and F420H2-dependent heterodisulfide reductase (HdrA2B2C2) in Methanosarcina acetivorans (2017), and Fd- and NADH-dependent ubiquinol reductase (FixABCX) in Azotobacter vinelandii (2017). The electron-bifurcating flavoprotein complexes known to date fall into four groups that have evolved independently, namely those containing EtfAB (CarED, LctCB, FixBA) with bound FAD, a NuoF homolog (HydB, HytB, or HylB) harboring FMN, NfnB with bound FAD, or HdrA harboring FAD. All these flavoproteins are cytoplasmic except for the membrane-associated protein FixABCX. The organisms-in which they have been found-are strictly anaerobic microorganisms except for the aerobe A. vinelandii. The electron-bifurcating complexes are involved in a variety of processes such as butyric acid fermentation, methanogenesis, acetogenesis, anaerobic lactate oxidation, dissimilatory sulfate reduction, anaerobic- dearomatization, nitrogen fixation, and CO2 fixation. They contribute to energy conservation via the energy-converting ferredoxin: NAD+ reductase complex Rnf or the energy-converting ferredoxin-dependent hydrogenase complex Ech. This Review describes how this mechanism was discovered.
Keywords: Ech-complex; Rnf-complex; crossed-over reduction potentials; electron bifurcation; electron-transferring flavoproteins (EtfAB); energy conservation; ferredoxin; flavodoxin.
Figures

References
-
- Aboulnaga E. H., Pinkenburg O., Schiffels J., El-Refai A., Buckel W., Selmer T. (2013). Effect of an oxygen-tolerant bifurcating butyryl coenzyme A dehydrogenase/electron-transferring flavoprotein complex from Clostridium difficile on butyrate production in Escherichia coli. J. Bacteriol. 195, 3704–3713. 10.1128/JB.00321-13 - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

