Pangenomic Definition of Prokaryotic Species and the Phylogenetic Structure of Prochlorococcus spp
- PMID: 29593678
- PMCID: PMC5857598
- DOI: 10.3389/fmicb.2018.00428
Pangenomic Definition of Prokaryotic Species and the Phylogenetic Structure of Prochlorococcus spp
Abstract
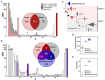
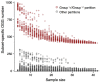
The pangenome is the collection of all groups of orthologous genes (OGGs) from a set of genomes. We apply the pangenome analysis to propose a definition of prokaryotic species based on identification of lineage-specific gene sets. While being similar to the classical biological definition based on allele flow, it does not rely on DNA similarity levels and does not require analysis of homologous recombination. Hence this definition is relatively objective and independent of arbitrary thresholds. A systematic analysis of 110 accepted species with the largest numbers of sequenced strains yields results largely consistent with the existing nomenclature. However, it has revealed that abundant marine cyanobacteria Prochlorococcus marinus should be divided into two species. As a control we have confirmed the paraphyletic origin of Yersinia pseudotuberculosis (with embedded, monophyletic Y. pestis) and Burkholderia pseudomallei (with B. mallei). We also demonstrate that by our definition and in accordance with recent studies Escherichia coli and Shigella spp. are one species.
Keywords: monophyly; pangenome; paraphyly; prokaryotic species; species definition; taxonomy.
Figures



References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

