Structure and Immune Recognition of the HIV Glycan Shield
- PMID: 29595997
- PMCID: PMC6163090
- DOI: 10.1146/annurev-biophys-060414-034156
Structure and Immune Recognition of the HIV Glycan Shield
Abstract
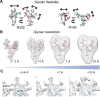
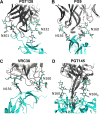
Vaccine design efforts against the human immunodeficiency virus (HIV) have been greatly stimulated by the observation that many infected patients eventually develop highly potent broadly neutralizing antibodies (bnAbs). Importantly, these bnAbs have evolved to recognize not only the two protein components of the viral envelope protein (Env) but also the numerous glycans that form a protective barrier on the Env protein. Because Env is heavily glycosylated compared to host glycoproteins, the glycans have become targets for the antibody response. Therefore, considerable efforts have been made in developing and validating biophysical methods to elucidate the complex structure of the Env-spike glycoprotein, with its combination of glycan and protein epitopes. We illustrate here how the application of robust biophysical methods has transformed our understanding of the structure and function of the HIV Env spike and stimulated innovation in vaccine design strategies that takes into account the essential glycan components.
Keywords: EM structure; X-ray structure; broadly neutralizing antibody; glycan composition; glycosylation; viral glycoprotein.
Figures

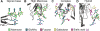
Similar articles
-
The HIV glycan shield as a target for broadly neutralizing antibodies.FEBS J. 2015 Dec;282(24):4679-91. doi: 10.1111/febs.13530. Epub 2015 Oct 23. FEBS J. 2015. PMID: 26411545 Free PMC article. Review.
-
Changes in Structure and Antigenicity of HIV-1 Env Trimers Resulting from Removal of a Conserved CD4 Binding Site-Proximal Glycan.J Virol. 2016 Sep 29;90(20):9224-36. doi: 10.1128/JVI.01116-16. Print 2016 Oct 15. J Virol. 2016. PMID: 27489265 Free PMC article.
-
Glycoengineering HIV-1 Env creates 'supercharged' and 'hybrid' glycans to increase neutralizing antibody potency, breadth and saturation.PLoS Pathog. 2018 May 2;14(5):e1007024. doi: 10.1371/journal.ppat.1007024. eCollection 2018 May. PLoS Pathog. 2018. PMID: 29718999 Free PMC article.
-
Comparative Analysis of the Glycosylation Profiles of Membrane-Anchored HIV-1 Envelope Glycoprotein Trimers and Soluble gp140.J Virol. 2015 Aug;89(16):8245-57. doi: 10.1128/JVI.00628-15. Epub 2015 May 27. J Virol. 2015. PMID: 26018173 Free PMC article.
-
Glycans in HIV-1 vaccine design - engaging the shield.Trends Microbiol. 2022 Sep;30(9):866-881. doi: 10.1016/j.tim.2022.02.004. Epub 2022 Mar 9. Trends Microbiol. 2022. PMID: 35279348 Review.
Cited by
-
Single-Molecule FRET Imaging of Virus Spike-Host Interactions.Viruses. 2021 Feb 21;13(2):332. doi: 10.3390/v13020332. Viruses. 2021. PMID: 33669922 Free PMC article. Review.
-
Structural basis of broad HIV neutralization by a vaccine-induced cow antibody.Sci Adv. 2020 May 27;6(22):eaba0468. doi: 10.1126/sciadv.aba0468. eCollection 2020 May. Sci Adv. 2020. PMID: 32518821 Free PMC article.
-
Networks of HIV-1 Envelope Glycans Maintain Antibody Epitopes in the Face of Glycan Additions and Deletions.Structure. 2020 Aug 4;28(8):897-909.e6. doi: 10.1016/j.str.2020.04.022. Epub 2020 May 19. Structure. 2020. PMID: 32433992 Free PMC article.
-
COVID-19 Vaccines: "Warp Speed" Needs Mind Melds, Not Warped Minds.J Virol. 2020 Aug 17;94(17):e01083-20. doi: 10.1128/JVI.01083-20. Print 2020 Aug 17. J Virol. 2020. PMID: 32591466 Free PMC article. Review.
-
Influence of glycosylation on the immunogenicity and antigenicity of viral immunogens.Biotechnol Adv. 2024 Jan-Feb;70:108283. doi: 10.1016/j.biotechadv.2023.108283. Epub 2023 Nov 14. Biotechnol Adv. 2024. PMID: 37972669 Free PMC article. Review.
References
-
- Agirre J, Davies GJ, Wilson KS, Cowtan KD. Carbohydrate structure: the rocky road to automation. Curr Opin Struct Biol. 2017;44:39–47. - PubMed
-
- Arnold JN, Radcliffe CM, Wormald MR, Royle L, Harvey DJ, et al. The glycosylation of human serum IgD and IgE and the accessibility of identified oligomannose structures for interaction with mannan-binding lectin. J Immunol. 2004;173:6831–40. - PubMed
-
- Baba TW, Liska V, Hofmann-Lehmann R, Vlasak J, Xu W, et al. Human neutralizing monoclonal antibodies of the IgG1 subtype protect against mucosal simian-human immunodeficiency virus infection. Nat Med. 2000;6:200–6. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

