The loss of SMG1 causes defects in quality control pathways in Physcomitrella patens
- PMID: 29596649
- PMCID: PMC6009662
- DOI: 10.1093/nar/gky225
The loss of SMG1 causes defects in quality control pathways in Physcomitrella patens
Abstract
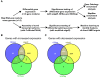
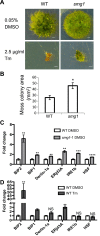
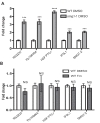
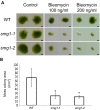
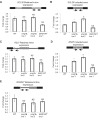
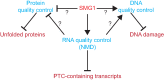
Nonsense-mediated mRNA decay (NMD) is important for RNA quality control and gene regulation in eukaryotes. NMD targets aberrant transcripts for decay and also directly influences the abundance of non-aberrant transcripts. In animals, the SMG1 kinase plays an essential role in NMD by phosphorylating the core NMD factor UPF1. Despite SMG1 being ubiquitous throughout the plant kingdom, little is known about its function, probably because SMG1 is atypically absent from the genome of the model plant, Arabidopsis thaliana. By combining our previously established SMG1 knockout in moss with transcriptome-wide analysis, we reveal the range of processes involving SMG1 in plants. Machine learning assisted analysis suggests that 32% of multi-isoform genes produce NMD-targeted transcripts and that splice junctions downstream of a stop codon act as the major determinant of NMD targeting. Furthermore, we suggest that SMG1 is involved in other quality control pathways, affecting DNA repair and the unfolded protein response, in addition to its role in mRNA quality control. Consistent with this, smg1 plants have increased susceptibility to DNA damage, but increased tolerance to unfolded protein inducing agents. The potential involvement of SMG1 in RNA, DNA and protein quality control has major implications for the study of these processes in plants.
Figures

Similar articles
-
SMG1 is an ancient nonsense-mediated mRNA decay effector.Plant J. 2013 Dec;76(5):800-10. doi: 10.1111/tpj.12329. Epub 2013 Nov 5. Plant J. 2013. PMID: 24103012
-
CK2-mediated TEL2 phosphorylation augments nonsense-mediated mRNA decay (NMD) by increase of SMG1 stability.Biochim Biophys Acta. 2013 Oct;1829(10):1047-55. doi: 10.1016/j.bbagrm.2013.06.002. Epub 2013 Jul 3. Biochim Biophys Acta. 2013. PMID: 23831331
-
MicroRNA 125 represses nonsense-mediated mRNA decay by regulating SMG1 expression.Biochem Biophys Res Commun. 2013 May 24;435(1):16-20. doi: 10.1016/j.bbrc.2013.03.129. Epub 2013 Apr 10. Biochem Biophys Res Commun. 2013. PMID: 23583196
-
Beyond quality control: The role of nonsense-mediated mRNA decay (NMD) in regulating gene expression.Semin Cell Dev Biol. 2018 Mar;75:78-87. doi: 10.1016/j.semcdb.2017.08.053. Epub 2017 Sep 1. Semin Cell Dev Biol. 2018. PMID: 28866327 Review.
-
The evolution and diversity of the nonsense-mediated mRNA decay pathway.F1000Res. 2018 Aug 15;7:1299. doi: 10.12688/f1000research.15872.2. eCollection 2018. F1000Res. 2018. PMID: 30345031 Free PMC article. Review.
Cited by
-
Direct RNA sequencing dataset of SMG1 KO mutant Physcomitrella (Physcomitrium patens).Data Brief. 2020 Nov 29;33:106602. doi: 10.1016/j.dib.2020.106602. eCollection 2020 Dec. Data Brief. 2020. PMID: 33313367 Free PMC article.
-
Distinct types of short open reading frames are translated in plant cells.Genome Res. 2019 Sep;29(9):1464-1477. doi: 10.1101/gr.253302.119. Epub 2019 Aug 6. Genome Res. 2019. PMID: 31387879 Free PMC article.
-
Widespread Exon Junction Complex Footprints in the RNA Degradome Mark mRNA Degradation before Steady State Translation.Plant Cell. 2020 Apr;32(4):904-922. doi: 10.1105/tpc.19.00666. Epub 2020 Jan 27. Plant Cell. 2020. PMID: 31988264 Free PMC article.
-
Quantitative proteomic dataset of the moss Physcomitrium patens SMG1 KO mutant line.Data Brief. 2021 Dec 11;40:107706. doi: 10.1016/j.dib.2021.107706. eCollection 2022 Feb. Data Brief. 2021. PMID: 34977295 Free PMC article.
-
Expression of a human cDNA in moss results in spliced mRNAs and fragmentary protein isoforms.Commun Biol. 2021 Aug 12;4(1):964. doi: 10.1038/s42003-021-02486-3. Commun Biol. 2021. PMID: 34385580 Free PMC article.
References
-
- Mühlemann O. Recognition of nonsense mRNA: towards a unified model. Biochem. Soc. Trans. 2008; 36:497–501. - PubMed
-
- He F., Li X., Spatrick P., Casillo R., Dong S., Jacobson A.. Genome-wide analysis of mRNAs regulated by the nonsense-mediated and 5′ to 3′ mRNA decay pathways in yeast. Mol. Cell. 2003; 12:1439–1452. - PubMed
-
- Mendell J.T., Sharifi N.A., Meyers J.L., Martinez-Murillo F., Dietz H.C.. Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise. Nat. Genet. 2004; 36:1073–1078. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

