Phylogenetic MLSA and phenotypic analysis identification of three probable novel Pseudomonas species isolated on King George Island, South Shetland, Antarctica
- PMID: 29598976
- PMCID: PMC6175711
- DOI: 10.1016/j.bjm.2018.02.005
Phylogenetic MLSA and phenotypic analysis identification of three probable novel Pseudomonas species isolated on King George Island, South Shetland, Antarctica
Abstract
Antarctica harbors a great diversity of microorganisms, including bacteria, archaea, microalgae and yeasts. The Pseudomonas genus is one of the most diverse and successful bacterial groups described to date, but only eight species isolated from Antarctica have been characterized. Here, we present three potentially novel species isolated on King George Island. The most abundant isolates from four different environments, were genotypically and phenotypically characterized. Multilocus sequence analysis and 16S rRNA gene analysis of a sequence concatenate for six genes (16S, aroE, glnS, gyrB, ileS and rpoD), determined one of the isolates to be a new Pseudomonas mandelii strain, while the other three are good candidates for new Pseudomonas species. Additionally, genotype analyses showed the three candidates to be part of a new subgroup within the Pseudomonas fluorescens complex, together with the Antarctic species Pseudomonas antarctica and Pseudomonas extremaustralis. We propose terming this new subgroup P. antarctica. Likewise, phenotypic analyses using API 20 NE and BIOLOG® corroborated the genotyping results, confirming that all presented isolates form part of the P. fluorescens complex. Pseudomonas genus research on the Antarctic continent is in its infancy. To understand these microorganisms' role in this extreme environment, the characterization and description of new species is vital.
Keywords: Antarctica; Multilocus Sequence Analysis; P. fluorescens complex; Pseudomonas.
Copyright © 2018 Sociedade Brasileira de Microbiologia. Published by Elsevier Editora Ltda. All rights reserved.
Figures

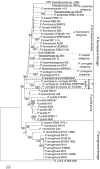
References
-
- Cowan D.A., Tow L.A. Endangered Antarctic environments. Annu Rev Microbiol. 2004;58:649–690. - PubMed
-
- Smith J., Tow L., Stafford W., Cary C., Cowan D. Bacterial diversity in three different Antarctic cold desert mineral soils. Microb Ecol. 2006;51(4):413–421. - PubMed
-
- Loperena L., Soria V., Varela H. Extracellular enzymes produced by microorganisms isolated from maritime Antarctica. World J Microbiol Biotechnol. 2012;28(5):2249–2256. - PubMed
-
- Cavicchioli R. Microbial ecology of Antarctic aquatic systems. Nat Rev Microbiol. 2015;13(11):691–706. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
