Genome-Wide Association Mapping Uncovers Fw1, a Dominant Gene Conferring Resistance to Fusarium Wilt in Strawberry
- PMID: 29602808
- PMCID: PMC5940171
- DOI: 10.1534/g3.118.200129
Genome-Wide Association Mapping Uncovers Fw1, a Dominant Gene Conferring Resistance to Fusarium Wilt in Strawberry
Abstract
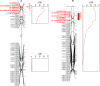
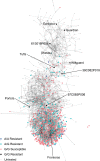
Fusarium wilt, a soil-borne disease caused by the fungal pathogen Fusarium oxysporum f. sp. fragariae, threatens strawberry (Fragaria × ananassa) production worldwide. The spread of the pathogen, coupled with disruptive changes in soil fumigation practices, have greatly increased disease pressure and the importance of developing resistant cultivars. While resistant and susceptible cultivars have been reported, a limited number of germplasm accessions have been analyzed, and contradictory conclusions have been reached in earlier studies to elucidate the underlying genetic basis of resistance. Here, we report the discovery of Fw1, a dominant gene conferring resistance to Fusarium wilt in strawberry. The Fw1 locus was uncovered in a genome-wide association study of 565 historically and commercially important strawberry accessions genotyped with 14,408 SNP markers. Fourteen SNPs in linkage disequilibrium with Fw1 physically mapped to a 2.3 Mb segment on chromosome 2 in a diploid F. vesca reference genome. Fw1 and 11 tightly linked GWAS-significant SNPs mapped to linkage group 2C in octoploid segregating populations. The most significant SNP explained 85% of the phenotypic variability and predicted resistance in 97% of the accessions tested-broad-sense heritability was 0.96. Several disease resistance and defense-related gene homologs, including a small cluster of genes encoding nucleotide-binding leucine-rich-repeat proteins, were identified in the 0.7 Mb genomic segment predicted to harbor Fw1 DNA variants and candidate genes identified in the present study should facilitate the development of high-throughput genotyping assays for accurately predicting Fusarium wilt phenotypes and applying marker-assisted selection.
Keywords: Fragaria; Fusarium wilt; innate immunity; polyploid; strawberry.
Copyright © 2018 Pincot et al.
Figures

Similar articles
-
Novel Fusarium wilt resistance genes uncovered in natural and cultivated strawberry populations are found on three non-homoeologous chromosomes.Theor Appl Genet. 2022 Jun;135(6):2121-2145. doi: 10.1007/s00122-022-04102-2. Epub 2022 May 18. Theor Appl Genet. 2022. PMID: 35583656 Free PMC article.
-
FaRXf1: a locus conferring resistance to angular leaf spot caused by Xanthomonas fragariae in octoploid strawberry.Theor Appl Genet. 2016 Jun;129(6):1191-201. doi: 10.1007/s00122-016-2695-1. Epub 2016 Feb 24. Theor Appl Genet. 2016. PMID: 26910360
-
A genome-wide association study uncovers consistent quantitative trait loci for resistance to Verticillium wilt and Fusarium wilt race 4 in the US Upland cotton.Theor Appl Genet. 2020 Feb;133(2):563-577. doi: 10.1007/s00122-019-03487-x. Epub 2019 Nov 25. Theor Appl Genet. 2020. PMID: 31768602
-
Secreted in Xylem 6 (SIX6) Mediates Fusarium oxysporum f. sp. fragariae Race 1 Avirulence on FW1-Resistant Strawberry Cultivars.Mol Plant Microbe Interact. 2024 Jun;37(6):530-541. doi: 10.1094/MPMI-02-24-0012-R. Epub 2024 Jun 24. Mol Plant Microbe Interact. 2024. PMID: 38552146
-
Characterization of Disease Resistance Loci in the USDA Soybean Germplasm Collection Using Genome-Wide Association Studies.Phytopathology. 2016 Oct;106(10):1139-1151. doi: 10.1094/PHYTO-01-16-0042-FI. Epub 2016 Jul 11. Phytopathology. 2016. PMID: 27135674 Review.
Cited by
-
Average semivariance yields accurate estimates of the fraction of marker-associated genetic variance and heritability in complex trait analyses.PLoS Genet. 2021 Aug 26;17(8):e1009762. doi: 10.1371/journal.pgen.1009762. eCollection 2021 Aug. PLoS Genet. 2021. PMID: 34437540 Free PMC article.
-
RNA interference-based strategies to control Botrytis cinerea infection in cultivated strawberry.Plant Cell Rep. 2024 Jul 24;43(8):201. doi: 10.1007/s00299-024-03288-7. Plant Cell Rep. 2024. PMID: 39048858 Free PMC article.
-
A roadmap for research in octoploid strawberry.Hortic Res. 2020 Mar 15;7:33. doi: 10.1038/s41438-020-0252-1. eCollection 2020. Hortic Res. 2020. PMID: 32194969 Free PMC article. Review.
-
A Dominance Hypothesis Argument for Historical Genetic Gains and the Fixation of Heterosis in Octoploid Strawberry.Genetics. 2024 Oct 10;228(4):iyae159. doi: 10.1093/genetics/iyae159. Online ahead of print. Genetics. 2024. PMID: 39385702 Free PMC article.
-
Disease Resistance Genetics and Genomics in Octoploid Strawberry.G3 (Bethesda). 2019 Oct 7;9(10):3315-3332. doi: 10.1534/g3.119.400597. G3 (Bethesda). 2019. PMID: 31420323 Free PMC article.
References
-
- Abdet-Sattar M., El-Marzoky H., Mohamed A., 2008. Occurrence of soilborne diseases and root knot nematodes in strawberry plants grown on compacted rice straw bales compared with naturally infested soils. J. Plant Prot. Res. 48(2): 223–235. 10.2478/v10045-008-0026-5 - DOI
-
- Alexander L. J., Tucker C. M., 1945. Physiologic specialization in the tomato wilt fungus Fusarium oxysporum f. sp. lycopersici. J. Agric. Res. 70: 303–313.
-
- The Arabidopsis Information Resource (TAIR), 2015 Locus: AT3G46530 Available at: http://www.arabidopsis.org/servlets/TairObject?id=35741&type=locus. Accessed: February 2, 2018.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
