Nuclease-free Adeno-Associated Virus-Mediated Il2rg Gene Editing in X-SCID Mice
- PMID: 29606506
- PMCID: PMC5993949
- DOI: 10.1016/j.ymthe.2018.02.028
Nuclease-free Adeno-Associated Virus-Mediated Il2rg Gene Editing in X-SCID Mice
Abstract
X-linked severe combined immunodeficiency (X-SCID) has been successfully treated by hematopoietic stem cell (HSC) transduction with retroviral vectors expressing the interleukin-2 receptor subunit gamma gene (IL2RG), but several patients developed malignancies due to vector integration near cellular oncogenes. This adverse side effect could in principle be avoided by accurate IL2RG gene editing with a vector that does not contain a functional promoter or IL2RG gene. Here, we show that adeno-associated virus (AAV) gene editing vectors can insert a partial Il2rg cDNA at the endogenous Il2rg locus in X-SCID murine bone marrow cells and that these ex vivo-edited cells repopulate transplant recipients and produce CD4+ and CD8+ T cells. Circulating, edited lymphocytes increased over time and appeared in secondary transplant recipients, demonstrating successful editing in long-term repopulating cells. Random vector integration events were nearly undetectable, and malignant transformation of the transplanted cells was not observed. Similar editing frequencies were observed in human hematopoietic cells. Our results demonstrate that therapeutically relevant HSC gene editing can be achieved by AAV vectors in the absence of site-specific nucleases and suggest that this may be a safe and effective therapy for hematopoietic diseases where in vivo selection can increase edited cell numbers.
Keywords: AAV vector; genome editing; hematopoietic stem cells.
Copyright © 2018 The American Society of Gene and Cell Therapy. Published by Elsevier Inc. All rights reserved.
Figures





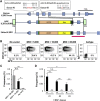
References
-
- Noguchi M., Yi H., Rosenblatt H.M., Filipovich A.H., Adelstein S., Modi W.S., McBride O.W., Leonard W.J. Interleukin-2 receptor gamma chain mutation results in X-linked severe combined immunodeficiency in humans. Cell. 1993;73:147–157. - PubMed
-
- Puck J.M., Deschênes S.M., Porter J.C., Dutra A.S., Brown C.J., Willard H.F., Henthorn P.S. The interleukin-2 receptor gamma chain maps to Xq13.1 and is mutated in X-linked severe combined immunodeficiency, SCIDX1. Hum. Mol. Genet. 1993;2:1099–1104. - PubMed
-
- Cavazzana-Calvo M., Hacein-Bey S., de Saint Basile G., Gross F., Yvon E., Nusbaum P., Selz F., Hue C., Certain S., Casanova J.L. Gene therapy of human severe combined immunodeficiency (SCID)-X1 disease. Science. 2000;288:669–672. - PubMed
-
- Gaspar H.B., Parsley K.L., Howe S., King D., Gilmour K.C., Sinclair J., Brouns G., Schmidt M., Von Kalle C., Barington T. Gene therapy of X-linked severe combined immunodeficiency by use of a pseudotyped gammaretroviral vector. Lancet. 2004;364:2181–2187. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

