Genome rearrangements and phylogeny reconstruction in Yersinia pestis
- PMID: 29607260
- PMCID: PMC5877447
- DOI: 10.7717/peerj.4545
Genome rearrangements and phylogeny reconstruction in Yersinia pestis
Abstract
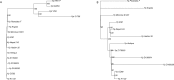
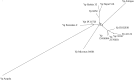
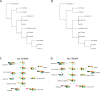
Genome rearrangements have played an important role in the evolution of Yersinia pestis from its progenitor Yersinia pseudotuberculosis. Traditional phylogenetic trees for Y. pestis based on sequence comparison have short internal branches and low bootstrap supports as only a small number of nucleotide substitutions have occurred. On the other hand, even a small number of genome rearrangements may resolve topological ambiguities in a phylogenetic tree. We reconstructed phylogenetic trees based on genome rearrangements using several popular approaches such as Maximum likelihood for Gene Order and the Bayesian model of genome rearrangements by inversions. We also reconciled phylogenetic trees for each of the three CRISPR loci to obtain an integrated scenario of the CRISPR cassette evolution. Analysis of contradictions between the obtained evolutionary trees yielded numerous parallel inversions and gain/loss events. Our data indicate that an integrated analysis of sequence-based and inversion-based trees enhances the resolution of phylogenetic reconstruction. In contrast, reconstructions of strain relationships based on solely CRISPR loci may not be reliable, as the history is obscured by large deletions, obliterating the order of spacer gains. Similarly, numerous parallel gene losses preclude reconstruction of phylogeny based on gene content.
Keywords: Bacteria evolution; Genome rearrangements; Phylogeny reconstruction.
Conflict of interest statement
Mikhail S. Gelfand is an Academic Editor for PeerJ.
Figures



Similar articles
-
Analysis of the three Yersinia pestis CRISPR loci provides new tools for phylogenetic studies and possibly for the investigation of ancient DNA.Adv Exp Med Biol. 2007;603:327-38. doi: 10.1007/978-0-387-72124-8_30. Adv Exp Med Biol. 2007. PMID: 17966429
-
Is homoplasy or lineage sorting the source of incongruent mtdna and nuclear gene trees in the stiff-tailed ducks (Nomonyx-Oxyura)?Syst Biol. 2005 Feb;54(1):35-55. doi: 10.1080/10635150590910249. Syst Biol. 2005. PMID: 15805009
-
Genomic comparison of Yersinia pestis and Yersinia pseudotuberculosis by combination of suppression subtractive hybridization and DNA microarray.Arch Microbiol. 2006 Aug;186(2):151-9. doi: 10.1007/s00203-006-0129-1. Epub 2006 Jul 11. Arch Microbiol. 2006. PMID: 16832628
-
Genome and Evolution of Yersinia pestis.Adv Exp Med Biol. 2016;918:171-192. doi: 10.1007/978-94-024-0890-4_6. Adv Exp Med Biol. 2016. PMID: 27722863 Review.
-
'Add, stir and reduce': Yersinia spp. as model bacteria for pathogen evolution.Nat Rev Microbiol. 2016 Mar;14(3):177-90. doi: 10.1038/nrmicro.2015.29. Nat Rev Microbiol. 2016. PMID: 26876035 Review.
Cited by
-
Genome rearrangements and selection in multi-chromosome bacteria Burkholderia spp.BMC Genomics. 2018 Dec 27;19(1):965. doi: 10.1186/s12864-018-5245-1. BMC Genomics. 2018. PMID: 30587126 Free PMC article.
-
Micro-evolution of three Streptococcus species: selection, antigenic variation, and horizontal gene inflow.BMC Evol Biol. 2019 Mar 27;19(1):83. doi: 10.1186/s12862-019-1403-6. BMC Evol Biol. 2019. PMID: 30917781 Free PMC article.
-
High Rates of Genome Rearrangements and Pathogenicity of Shigella spp.Front Microbiol. 2021 Apr 12;12:628622. doi: 10.3389/fmicb.2021.628622. eCollection 2021. Front Microbiol. 2021. PMID: 33912145 Free PMC article.
-
Genomic structural plasticity of rodent-associated Bartonella in nature.Mol Ecol. 2022 Jul;31(14):3784-3797. doi: 10.1111/mec.16547. Epub 2022 Jun 10. Mol Ecol. 2022. PMID: 35620948 Free PMC article.
References
-
- Achtman M, Zurth K, Morelli G, Torrea G, Guiyoule A, Carnie E. Yersinia pestis, the cause of plague, is a recently emerged clone of Yersinia pseudotuberculosis. Proceedings of the National Academy of Sciences of the United States of America. 1999;96(24):14043–14048. doi: 10.1073/pnas.96.24.14043. - DOI - PMC - PubMed
-
- Chain PSG, Hu P, Malfatti SA, Radnedge L, Larimer F, Vergez LM, Worsham P, Chu MC, Andersen GL. Complete genome sequence of Yersinia pestis strains Antiqua and Nepal516: evidence of gene reduction in an emerging pathogen. Journal of Bacteriology. 2006;188(12):4453–4463. doi: 10.1128/JB.00124-06. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

