Homologous Recombination between Genetically Divergent Campylobacter fetus Lineages Supports Host-Associated Speciation
- PMID: 29608720
- PMCID: PMC5830970
- DOI: 10.1093/gbe/evy048
Homologous Recombination between Genetically Divergent Campylobacter fetus Lineages Supports Host-Associated Speciation
Abstract
Homologous recombination is a major driver of bacterial speciation. Genetic divergence and host association are important factors influencing homologous recombination. Here, we study these factors for Campylobacter fetus, which shows a distinct intraspecific host dichotomy. Campylobacter fetus subspecies fetus (Cff) and venerealis are associated with mammals, whereas C. fetus subsp. testudinum (Cft) is associated with reptiles. Recombination between these genetically divergent C. fetus lineages is extremely rare. Previously it was impossible to show whether this barrier to recombination was determined by the differential host preferences, by the genetic divergence between both lineages or by other factors influencing recombination, such as restriction-modification, CRISPR/Cas, and transformation systems. Fortuitously, a distinct C. fetus lineage (ST69) was found, which was highly related to mammal-associated C. fetus, yet isolated from a chelonian. The whole genome sequences of two C. fetus ST69 isolates were compared with those of mammal- and reptile-associated C. fetus strains for phylogenetic and recombination analysis. In total, 5.1-5.5% of the core genome of both ST69 isolates showed signs of recombination. Of the predicted recombination regions, 80.4% were most closely related to Cft, 14.3% to Cff, and 5.6% to C. iguaniorum. Recombination from C. fetus ST69 to Cft was also detected, but to a lesser extent and only in chelonian-associated Cft strains. This study shows that despite substantial genetic divergence no absolute barrier to homologous recombination exists between two distinct C. fetus lineages when occurring in the same host type, which provides valuable insights in bacterial speciation and evolution.
Figures
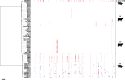

Similar articles
-
Comparative Genomics of Campylobacter fetus from Reptiles and Mammals Reveals Divergent Evolution in Host-Associated Lineages.Genome Biol Evol. 2016 Jul 2;8(6):2006-19. doi: 10.1093/gbe/evw146. Genome Biol Evol. 2016. PMID: 27333878 Free PMC article.
-
Genetic divergence of Campylobacter fetus strains of mammal and reptile origins.J Clin Microbiol. 2005 Jul;43(7):3334-40. doi: 10.1128/JCM.43.7.3334-3340.2005. J Clin Microbiol. 2005. PMID: 16000457 Free PMC article.
-
Assessing the intra-species genetic variability in the clonal pathogen Campylobacter fetus: CRISPRs are highly polymorphic DNA markers.J Microbiol Methods. 2017 Jan;132:86-94. doi: 10.1016/j.mimet.2016.11.012. Epub 2016 Nov 17. J Microbiol Methods. 2017. PMID: 27867047
-
[Pathogenesis of Campylobacter fetus infection].Fukuoka Igaku Zasshi. 2003 Jul;94(7):225-34. Fukuoka Igaku Zasshi. 2003. PMID: 14509230 Review. Japanese.
-
New molecular microbiology approaches in the study of Campylobacter fetus.Microb Biotechnol. 2011 Jan;4(1):8-19. doi: 10.1111/j.1751-7915.2010.00173.x. Microb Biotechnol. 2011. PMID: 21255368 Free PMC article. Review.
Cited by
-
L-cysteine transporter-PCR to detect hydrogen sulfide-producing Campylobacter fetus.PeerJ. 2019 Nov 5;7:e7820. doi: 10.7717/peerj.7820. eCollection 2019. PeerJ. 2019. PMID: 31720099 Free PMC article.
-
Genomic insights into the diversity, antimicrobial resistance and zoonotic potential of Campylobacter fetus across diverse hosts and geographies.Microb Genom. 2025 Jul;11(7):001446. doi: 10.1099/mgen.0.001446. Microb Genom. 2025. PMID: 40638214 Free PMC article.
-
Pathogenomics of Emerging Campylobacter Species.Clin Microbiol Rev. 2019 Jul 3;32(4):e00072-18. doi: 10.1128/CMR.00072-18. Print 2019 Sep 18. Clin Microbiol Rev. 2019. PMID: 31270126 Free PMC article. Review.
-
Living in Cold Blood: Arcobacter, Campylobacter, and Helicobacter in Reptiles.Front Microbiol. 2019 May 15;10:1086. doi: 10.3389/fmicb.2019.01086. eCollection 2019. Front Microbiol. 2019. PMID: 31191467 Free PMC article. Review.
-
Phylogenomic Analysis of Campylobacter fetus Reveals a Clonal Structure of Insertion Element ISCfe1 Positive Genomes.Front Microbiol. 2020 Nov 12;11:585374. doi: 10.3389/fmicb.2020.585374. eCollection 2020. Front Microbiol. 2020. PMID: 33281781 Free PMC article.
References
-
- Blaser MJ, Newell DG, Thompson SA, Zechner EL. (2008). Pathogenesis of Campylobacter fetus In: Nachamkin I, Szymanski CM, Blaser MJ, editors. Campylobacter. Washington (DC: ): ASM Press; p. 401–428.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

