Conservation and Divergence in the Candida Species Biofilm Matrix Mannan-Glucan Complex Structure, Function, and Genetic Control
- PMID: 29615504
- PMCID: PMC5885036
- DOI: 10.1128/mBio.00451-18
Conservation and Divergence in the Candida Species Biofilm Matrix Mannan-Glucan Complex Structure, Function, and Genetic Control
Abstract
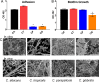
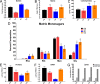
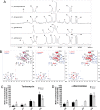
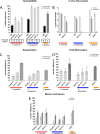
Candida biofilms resist the effects of available antifungal therapies. Prior studies with Candida albicans biofilms show that an extracellular matrix mannan-glucan complex (MGCx) contributes to antifungal sequestration, leading to drug resistance. Here we implement biochemical, pharmacological, and genetic approaches to explore a similar mechanism of resistance for the three most common clinically encountered non-albicansCandida species (NAC). Our findings reveal that each Candida species biofilm synthesizes a mannan-glucan complex and that the antifungal-protective function of this complex is conserved. Structural similarities extended primarily to the polysaccharide backbone (α-1,6-mannan and β-1,6-glucan). Surprisingly, biochemical analysis uncovered stark differences in the branching side chains of the MGCx among the species. Consistent with the structural analysis, similarities in the genetic control of MGCx production for each Candida species also appeared limited to the synthesis of the polysaccharide backbone. Each species appears to employ a unique subset of modification enzymes for MGCx synthesis, likely accounting for the observed side chain diversity. Our results argue for the conservation of matrix function among Candida spp. While biogenesis is preserved at the level of the mannan-glucan complex backbone, divergence emerges for construction of branching side chains. Thus, the MGCx backbone represents an ideal drug target for effective pan-Candida species biofilm therapy.IMPORTANCECandida species, the most common fungal pathogens, frequently grow as a biofilm. These adherent communities tolerate extremely high concentrations of antifungal agents, due in large part, to a protective extracellular matrix. The present studies define the structural, functional, and genetic similarities and differences in the biofilm matrix from the four most common Candida species. Each species synthesizes an extracellular mannan-glucan complex (MGCx) which contributes to sequestration of antifungal drug, shielding the fungus from this external assault. Synthesis of a common polysaccharide backbone appears conserved. However, subtle structural differences in the branching side chains likely rely upon unique modification enzymes, which are species specific. Our findings identify MGCx backbone synthesis as a potential pan-Candida biofilm therapeutic target.
Keywords: Candida; antifungal resistance; biofilm; extracellular matrix; non-albicans.
Copyright © 2018 Dominguez et al.
Figures


Similar articles
-
Conserved Role for Biofilm Matrix Polysaccharides in Candida auris Drug Resistance.mSphere. 2019 Jan 2;4(1):e00680-18. doi: 10.1128/mSphereDirect.00680-18. mSphere. 2019. PMID: 30602527 Free PMC article.
-
Role of the extracellular matrix in Candida biofilm antifungal resistance.FEMS Microbiol Rev. 2023 Nov 1;47(6):fuad059. doi: 10.1093/femsre/fuad059. FEMS Microbiol Rev. 2023. PMID: 37816666
-
Novel entries in a fungal biofilm matrix encyclopedia.mBio. 2014 Aug 5;5(4):e01333-14. doi: 10.1128/mBio.01333-14. mBio. 2014. PMID: 25096878 Free PMC article.
-
Mechanisms of Candida biofilm drug resistance.Future Microbiol. 2013 Oct;8(10):1325-37. doi: 10.2217/fmb.13.101. Future Microbiol. 2013. PMID: 24059922 Free PMC article. Review.
-
Fungal biofilms and antimycotics.Curr Drug Targets. 2005 Dec;6(8):887-94. doi: 10.2174/138945005774912762. Curr Drug Targets. 2005. PMID: 16375672 Review.
Cited by
-
Natural Anti-biofilm Agents: Strategies to Control Biofilm-Forming Pathogens.Front Microbiol. 2020 Oct 29;11:566325. doi: 10.3389/fmicb.2020.566325. eCollection 2020. Front Microbiol. 2020. PMID: 33193155 Free PMC article. Review.
-
Two promising Bacillus-derived antifungal lipopeptide leads AF4 and AF5 and their combined effect with fluconazole on the in vitro Candida glabrata biofilms.Front Pharmacol. 2024 Apr 19;15:1334419. doi: 10.3389/fphar.2024.1334419. eCollection 2024. Front Pharmacol. 2024. PMID: 38708082 Free PMC article.
-
Lactobacillus crispatus BC1 Biosurfactant Delivered by Hyalurosomes: An Advanced Strategy to Counteract Candida Biofilm.Antibiotics (Basel). 2021 Jan 1;10(1):33. doi: 10.3390/antibiotics10010033. Antibiotics (Basel). 2021. PMID: 33401413 Free PMC article.
-
Contribution of N-Linked Mannosylation Pathway to Candida parapsilosis and Candida tropicalis Biofilm Formation.Infect Drug Resist. 2023 Oct 26;16:6843-6857. doi: 10.2147/IDR.S431745. eCollection 2023. Infect Drug Resist. 2023. PMID: 37908782 Free PMC article.
-
Mechanisms of pathogenicity for the emerging fungus Candida auris.PLoS Pathog. 2023 Dec 21;19(12):e1011843. doi: 10.1371/journal.ppat.1011843. eCollection 2023 Dec. PLoS Pathog. 2023. PMID: 38127686 Free PMC article. Review.
References
-
- Pfaller MA, Andes DR, Diekema DJ, Horn DL, Reboli AC, Rotstein C, Franks B, Azie NE. 2014. Epidemiology and outcomes of invasive candidiasis due to non-albicans species of Candida in 2,496 patients: data from the Prospective Antifungal Therapy (PATH) registry 2004-2008. PLoS One 9:e101510. doi:10.1371/journal.pone.0101510. - DOI - PMC - PubMed
-
- Cleveland AA, Farley MM, Harrison LH, Stein B, Hollick R, Lockhart SR, Magill SS, Derado G, Park BJ, Chiller TM. 2012. Changes in incidence and antifungal drug resistance in candidemia: results from population-based laboratory surveillance in Atlanta and Baltimore, 2008-2011. Clin Infect Dis 55:1352–1361. doi:10.1093/cid/cis697. - DOI - PMC - PubMed
-
- Lockhart SR, Iqbal N, Cleveland AA, Farley MM, Harrison LH, Bolden CB, Baughman W, Stein B, Hollick R, Park BJ, Chiller T. 2012. Species identification and antifungal susceptibility testing of Candida bloodstream isolates from population-based surveillance studies in two U.S. cities from 2008 to 2011. J Clin Microbiol 50:3435–3442. doi:10.1128/JCM.01283-12. - DOI - PMC - PubMed
-
- Andes DR, Safdar N, Baddley JW, Alexander B, Brumble L, Freifeld A, Hadley S, Herwaldt L, Kauffman C, Lyon GM, Morrison V, Patterson T, Perl T, Walker R, Hess T, Chiller T, Pappas PG, The TRANSNET Investigators . 2016. The epidemiology and outcomes of invasive Candida infections among organ transplant recipients in the United States: results of the Transplant-Associated Infection Surveillance Network (TRANSNET). Transpl Infect Dis 18:921–931. doi:10.1111/tid.12613. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

