Fingerprinting Non-Terran Biosignatures
- PMID: 29634318
- PMCID: PMC6067094
- DOI: 10.1089/ast.2017.1712
Fingerprinting Non-Terran Biosignatures
Abstract
Most strategies for life detection rely upon finding features known to be associated with terran life, such as particular classes of molecules. But life may be vastly different on other planets and moons, particularly as we expand our efforts to explore ocean worlds like Europa and Enceladus. We propose a new concept for life detection that harnesses the power of DNA sequencing to yield intricate informatics fingerprints, even for life that is not nucleic acid-based. The concept is based on the fact that folded nucleic acid structures (aptamers) have been shown to be capable of binding a wide variety of compounds, whether inorganic, organic, or polymeric, and irrespective of being from a biotic or abiotic source. Each nucleic acid sequence can be thought of as a code, and a combination of codes as a "fingerprint." Over multiple analytes, the "fingerprint" of a non-terran sample can be analyzed by chemometric protocols to provide a classifier of molecular patterns and complexity. Ultimately the chemometric fingerprints of living systems, which may differ significantly from nonliving systems, could provide an empirical, agnostic means of detecting life. Because nucleic acids are exponentially amplified by the polymerase chain reaction, even very small input signals could be translated into a robust readable output. The derived sequences could be identified by a small, portable sequencing device or by capture and optical imaging on a DNA microarray. Without presupposing any particular molecular framework, this agnostic approach to life detection could be used from Mars to the far reaches of the Solar System, all within the framework of an instrument drawing little heat and power. Key Words: Agnostic biosignatures-Astrobiology-Chemometrics-DNA sequencing-Life detection-Proximity ligation assay. Astrobiology 18, 915-922.
Conflict of interest statement
No competing financial interests exist.
Figures


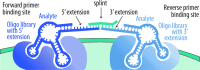
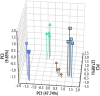
References
-
- Abraham A. (2005) Artificial neural networks. In Handbook of Measuring System Design, Part 8. Elements: B—Signal Conditioning, Wiley, Chichester, UK, doi: 10.1002/0471497398.mm421 - DOI
-
- Anders E. (1989) Pre-biotic organic matter from comets and asteroids. Nature 342:255–257 - PubMed
-
- Benardini J.N., III, La Duc M.T., Beaudet R.A., and Koukol R. (2014) Implementing planetary protection measures on the Mars Science Laboratory. Astrobiology 14:27–32 - PubMed
-
- Bhagat A.A.S., Bow H., Hou H.W., Tan S.J., Han J., and Lim C.T. (2010) Microfluidics for cell separation. Med Biol Eng Comput 48:999–1014 - PubMed
-
- Bywaters K.B., Schmidt H., Vercoutere W., Deamer D., Hawkins A.R., Quinn R.C., Burton A.S., and McKay C.P. (2017) Development of solid-state nanopore technology for life detection [abstract 3437]. In Astrbiology Science Conference Abstracts, Lunar and Planetary Institute, Houston
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

