A guide to chemokines and their receptors
- PMID: 29637711
- PMCID: PMC6120486
- DOI: 10.1111/febs.14466
A guide to chemokines and their receptors
Abstract
The chemokines (or chemotactic cytokines) are a large family of small, secreted proteins that signal through cell surface G protein-coupled heptahelical chemokine receptors. They are best known for their ability to stimulate the migration of cells, most notably white blood cells (leukocytes). Consequently, chemokines play a central role in the development and homeostasis of the immune system, and are involved in all protective or destructive immune and inflammatory responses. Classically viewed as inducers of directed chemotactic migration, it is now clear that chemokines can stimulate a variety of other types of directed and undirected migratory behavior, such as haptotaxis, chemokinesis, and haptokinesis, in addition to inducing cell arrest or adhesion. However, chemokine receptors on leukocytes can do more than just direct migration, and these molecules can also be expressed on, and regulate the biology of, many nonleukocytic cell types. Chemokines are profoundly affected by post-translational modification, by interaction with the extracellular matrix (ECM), and by binding to heptahelical 'atypical' chemokine receptors that regulate chemokine localization and abundance. This guide gives a broad overview of the chemokine and chemokine receptor families; summarizes the complex physical interactions that occur in the chemokine network; and, using specific examples, discusses general principles of chemokine function, focusing particularly on their ability to direct leukocyte migration.
Keywords: atypical chemokine receptor; cell migration; chemokine; chemokine receptor; glycosaminoglycan; immune surveillance; inflammation; leukocyte; oligomerization; protease.
© 2018 The Authors. The FEBS Journal published by John Wiley & Sons Ltd on behalf of Federation of European Biochemical Societies.
Figures
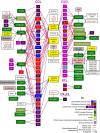
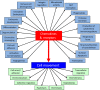


References
-
- Gonzalez E, Kulkarni H, Bolivar H, Mangano A, Sanchez R, Catano G, Nibbs RJ, Freedman BI, Quinones MP, Bamshad MJ et al (2005) The influence of CCL3L1 gene‐containing segmental duplications on HIV‐1/AIDS susceptibility. Science 307, 1434–1440. - PubMed
-
- Townson JR, Barcellos LF & Nibbs RJB (2002) Gene copy number regulates the production of the human chemokine CCL3‐L1. Eur J Immunol 32, 3016–3026. - PubMed
-
- Nakano H & Gunn MD (2001) Gene duplications at the chemokine locus on mouse chromosome 4: multiple strain‐specific haplotypes and the deletion of secondary lymphoid‐organ chemokine and EBI‐1 ligand chemokine genes in the plt mutation. J Immunol 166, 361–369. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

