A Cost-Effective High-Throughput Plasma and Serum Proteomics Workflow Enables Mapping of the Molecular Impact of Total Pancreatectomy with Islet Autotransplantation
- PMID: 29641209
- PMCID: PMC6791589
- DOI: 10.1021/acs.jproteome.8b00111
A Cost-Effective High-Throughput Plasma and Serum Proteomics Workflow Enables Mapping of the Molecular Impact of Total Pancreatectomy with Islet Autotransplantation
Abstract
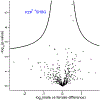
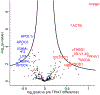
Blood is an ideal body fluid for the discovery or monitoring of diagnostic and prognostic protein biomarkers. However, discovering robust biomarkers requires the analysis of large numbers of samples to appropriately represent interindividual variability. To address this analytical challenge, we established a high-throughput and cost-effective proteomics workflow for accurate and comprehensive proteomics at an analytical depth applicable for clinical studies. For validation, we processed 1 μL each from 62 plasma samples in 96-well plates and analyzed the product by quantitative data-independent acquisition liquid chromatography/mass spectrometry; the data were queried using feature quantification with Spectronaut. To show the applicability of our workflow to serum, we analyzed a unique set of samples from 48 chronic pancreatitis patients, pre and post total pancreatectomy with islet autotransplantation (TPIAT) surgery. We identified 16 serum proteins with statistically significant abundance alterations, which represent a molecular signature distinct from that of chronic pancreatitis. In summary, we established a cost-efficient high-throughput workflow for comprehensive proteomics using PVDF-membrane-based digestion that is robust, automatable, and applicable to small plasma and serum volumes, e.g., finger stick. Application of this plasma/serum proteomics workflow resulted in the first mapping of the molecular implications of TPIAT on the serum proteome.
Keywords: TPIAT; biomarker; data-independent acquisition; plasma; serum.
Conflict of interest statement
Conflict of Interest Pertaining to the Presented Work:
VA receives compensation for consultancy and for being a member of an advisory board for Merck (MSD) and Janssen. The other authors declare no competing interests.
Figures
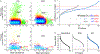

References
-
- Malekzadeh A; Twaalfhoven H; Wijnstok NJ; Killestein J; Blankenstein MA; Teunissen CE Comparison of Multiplex Platforms for Cytokine Assessments and Their Potential Use for Biomarker Profiling in Multiple Sclerosis. Cytokine 2017, 91, 145–152. - PubMed
-
- Mann M Origins of Mass Spectrometry-Based Proteomics. Nat. Rev. Mol. Cell Biol. 2016, 17 (11), 678–678. - PubMed
-
- Anderson NL The Human Plasma Proteome: History, Character, and Diagnostic Prospects. Mol. Cell. Proteomics 2002, 1 (11), 845–867. - PubMed
-
- Geyer PE; Kulak NA; Pichler G; Holdt LM; Teupser D; Mann M Plasma Proteome Profiling to Assess Human Health and Disease. Cell Syst. 2016, 2 (3), 185–195. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

