A role for the CXCR4-CXCL12 axis in the little skate, Leucoraja erinacea
- PMID: 29641231
- PMCID: PMC6139610
- DOI: 10.1152/ajpregu.00322.2017
A role for the CXCR4-CXCL12 axis in the little skate, Leucoraja erinacea
Abstract
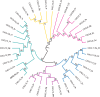
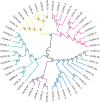
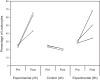
The interaction between C-X-C chemokine receptor type 4 (CXCR4) and its cognate ligand C-X-C motif chemokine ligand 12 (CXCL12) plays a critical role in regulating hematopoietic stem cell activation and subsequent cellular mobilization. Extensive studies of these genes have been conducted in mammals, but much less is known about the expression and function of CXCR4 and CXCL12 in non-mammalian vertebrates. In the present study, we identify simultaneous expression of CXCR4 and CXCL12 orthologs in the epigonal organ (the primary hematopoietic tissue) of the little skate, Leucoraja erinacea. Genetic and phylogenetic analyses were functionally supported by significant mobilization of leukocytes following administration of Plerixafor, a CXCR4 antagonist and clinically important drug. Our results provide evidence that, as in humans, Plerixafor disrupts CXCR4/CXCL12 binding in the little skate, facilitating release of leukocytes into the bloodstream. Our study illustrates the value of the little skate as a model organism, particularly in studies of hematopoiesis and potentially for preclinical research on hematological and vascular disorders.
Keywords: C-X-C chemokine ligand 12; C-X-C chemokine receptor type 4; Plerixafor/AMD3100; elasmobranch; mobilization.
Figures


References
-
- Bajoghli B, Aghaallaei N, Hess I, Rode I, Netuschil N, Tay BH, Venkatesh B, Yu JK, Kaltenbach SL, Holland ND, Diekhoff D, Happe C, Schorpp M, Boehm T. Evolution of genetic networks underlying the emergence of thymopoiesis in vertebrates. Cell 138: 186–197, 2009. doi: 10.1016/j.cell.2009.04.017. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

