Kaposi's Sarcoma-Associated Herpesvirus mRNA Accumulation in Nuclear Foci Is Influenced by Viral DNA Replication and Viral Noncoding Polyadenylated Nuclear RNA
- PMID: 29643239
- PMCID: PMC6002709
- DOI: 10.1128/JVI.00220-18
Kaposi's Sarcoma-Associated Herpesvirus mRNA Accumulation in Nuclear Foci Is Influenced by Viral DNA Replication and Viral Noncoding Polyadenylated Nuclear RNA
Abstract
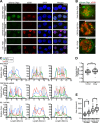
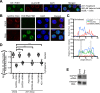
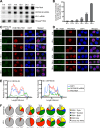
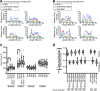
Kaposi's sarcoma-associated herpesvirus (KSHV), like other herpesviruses, replicates within the nuclei of its human cell host and hijacks host machinery for expression of its genes. The activities that culminate in viral DNA synthesis and assembly of viral proteins into capsids physically concentrate in nuclear areas termed viral replication compartments. We sought to better understand the spatiotemporal regulation of viral RNAs during the KSHV lytic phase by examining and quantifying the subcellular localization of select viral transcripts. We found that viral mRNAs, as expected, localized to the cytoplasm throughout the lytic phase. However, dependent on active viral DNA replication, viral transcripts also accumulated in the nucleus, often in foci in and around replication compartments, independent of the host shutoff effect. Our data point to involvement of the viral long noncoding polyadenylated nuclear (PAN) RNA in the localization of an early, intronless viral mRNA encoding ORF59-58 to nuclear foci that are associated with replication compartments.IMPORTANCE Late in the lytic phase, mRNAs from Kaposi's sarcoma-associated herpesvirus accumulate in the host cell nucleus near viral replication compartments, centers of viral DNA synthesis and virion production. This work contributes spatiotemporal data on herpesviral mRNAs within the lytic host cell and suggests a mechanism for viral RNA accumulation. Our findings indicate that the mechanism is independent of the host shutoff effect and splicing but dependent on active viral DNA synthesis and in part on the viral noncoding RNA, PAN RNA. PAN RNA is essential for the viral life cycle, and its contribution to the nuclear accumulation of viral messages may facilitate propagation of the virus.
Keywords: DNA replication compartments; KSHV; mRNA localization; nuclear foci; viral noncoding PAN RNA.
Copyright © 2018 American Society for Microbiology.
Figures


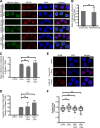
References
-
- Arias C, Weisburd B, Stern-Ginossar N, Mercier A, Madrid AS, Bellare P, Holdorf M, Weissman JS, Ganem D. 2014. KSHV 2.0: a comprehensive annotation of the Kaposi's sarcoma-associated herpesvirus genome using next-generation sequencing reveals novel genomic and functional features. PLoS Pathog 10:e1003847. doi: 10.1371/journal.ppat.1003847. - DOI - PMC - PubMed
-
- Pfeffer S, Sewer A, Lagos-Quintana M, Sheridan R, Sander C, Grasser FA, van Dyk LF, Ho CK, Shuman S, Chien M, Russo JJ, Ju J, Randall G, Lindenbach BD, Rice CM, Simon V, Ho DD, Zavolan M, Tuschl T. 2005. Identification of microRNAs of the herpesvirus family. Nat Methods 2:269–276. doi: 10.1038/nmeth746. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

